Vector Control: Indoor Residual Spraying
VectorControl_IRS.Rmd
# Load the requisite packages:
library(malariasimulation)
library(malariaEquilibrium)
# Set colour palette:
cols <- c("#E69F00", "#56B4E9", "#009E73", "#F0E442", "#0072B2", "#D55E00", "#CC79A7")Indoor Residual Spraying (IRS) involves periodically treating indoor
walls with insecticides to eliminate adult female mosquitoes that rest
indoors. malariasimulation can be used to investigate the
effect of malaria control strategies that deploy IRS. Users can set IRS
in the model using the set_spraying() function to
parameterise the time and coverage of spraying campaigns.
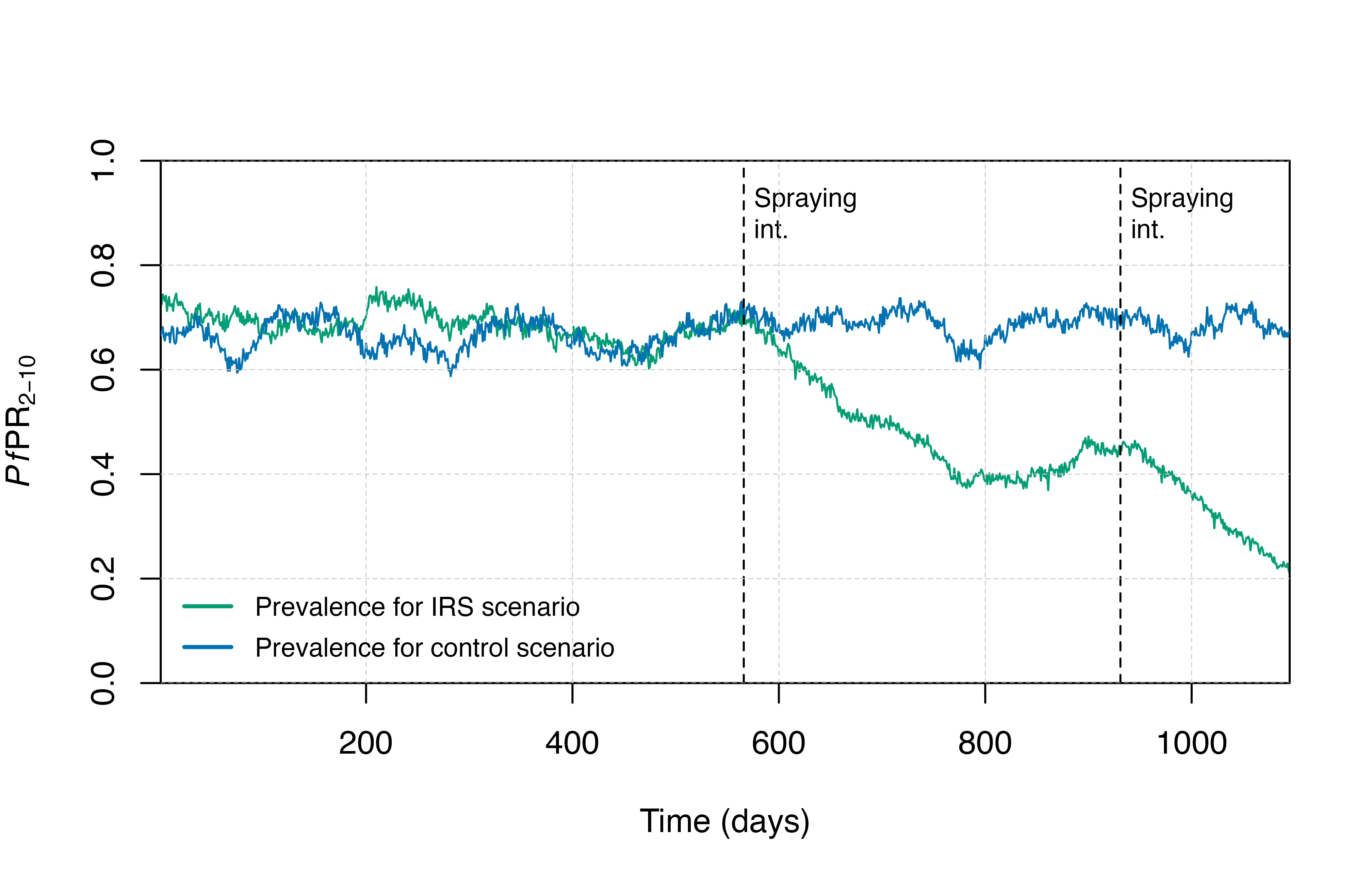
We will create a few plotting functions to visualise the output.
# Plotting functions
plot_prev <- function() {
plot(x = output$timestep, y = output$n_detect_lm_730_3650 / output$n_age_730_3650,
type = "l", col = cols[3], lwd = 1,
xlab = "Time (days)", ylab = expression(paste(italic(Pf),"PR"[2-10])),
xaxs = "i", yaxs = "i", ylim = c(0,1))
lines(x = output_control$timestep, y = output_control$n_detect_lm_730_3650 / output_control$n_age_730_3650,
col = cols[5], lwd = 1)
abline(v = sprayingtimesteps, lty = 2, lwd = 1, col = "black")
text(x = sprayingtimesteps + 10, y = 0.9, labels = "Spraying\nint.", adj = 0, cex = 0.8)
grid(lty = 2, col = "grey80", lwd = 0.5)
legend("bottomleft", box.lty = 0,
legend = c("Prevalence for IRS scenario","Prevalence for control scenario"),
col = c(cols[3], cols[5]), lty = c(1,1), lwd = 2, cex = 0.8, y.intersp = 1.3)
}Setting IRS parameters
Parameterisation
Use the get_parameters() function to generate a list of
parameters, accepting most of the default values, but modifying
seasonality values to model a seasonal setting. Then, we use the
set_equilibrium() function to to initialise the model at a
given entomological innoculation rate (EIR).
year <- 365
month <- 30
sim_length <- 3 * year
human_population <- 1000
starting_EIR <- 50
simparams <- get_parameters(
list(
human_population = human_population,
# seasonality parameters
model_seasonality = TRUE,
g0 = 0.285277,
g = c(-0.0248801, -0.0529426, -0.0168910),
h = c(-0.0216681, -0.0242904, -0.0073646)
)
)
simparams <- set_equilibrium(parameters = simparams, init_EIR = starting_EIR)
# Running simulation with no IRS
output_control <- run_simulation(timesteps = sim_length, parameters = simparams)A note on mosquito species
It is also possible to use the set_species() function to
account for 3 different mosquito species in the simulation. In this
case, the matrices would need to have additional column corresponding to
each mosquito species. For example, if we specified that there were 3
species of mosquitoes in the model and nets were distributed at two
timesteps, then the matrices would have 2 rows and 3 columns. If you are
not already familiar with the set_species() function, see
the Mosquito
Species vignette.
The default parameters are set to model Anopheles gambiae.
simparams$species
#> [1] "gamb"
simparams$species_proportions
#> [1] 1Simulation
Then we can run the simulations for a variety of IRS strategies. In
the example below, there are two rounds of IRS, the first at 30%
coverage and the second at 80% coverage, each 3 months prior to peak
rainfall for that year. The function peak_season_offset()
outputs a timestep when peak rainfall will be observed based on the
seasonality profile we set above. Notice that the matrices specified for
the parameters for ls_theta, ls_gamma,
ks_theta, ks_gamma, ms_theta, and
ms_gamma have two rows, one for each timestep where IRS is
implemented, and a number of columns corresponding to mosquito species.
In this example, we only have 1 column because the species is set to
“gamb” as we saw above.
The structure for the IRS model is documented in the supplementary information from Table 3 in the Supplementary Information of Sherrard-Smith et al., 2018. Table S2.1 in the Supplementary Appendix of Sherrard-Smith et al., 2022 has parameter estimates for insecticide resistance for IRS.
peak <- peak_season_offset(simparams)
sprayingtimesteps <- c(1, 2) * year + peak - 3 * month # A round of IRS is implemented in the 1st and second year 3 months prior to peak transmission.
sprayingparams <- set_spraying(
simparams,
timesteps = sprayingtimesteps,
coverages = c(0.3, 0.8), # # The first round covers 30% of the population and the second covers 80%.
ls_theta = matrix(2.025, nrow=length(sprayingtimesteps), ncol=1), # Matrix of mortality parameters; nrows=length(timesteps), ncols=length(species)
ls_gamma = matrix(-0.009, nrow=length(sprayingtimesteps), ncol=1), # Matrix of mortality parameters per round of IRS and per species
ks_theta = matrix(-2.222, nrow=length(sprayingtimesteps), ncol=1), # Matrix of feeding success parameters per round of IRS and per species
ks_gamma = matrix(0.008, nrow=length(sprayingtimesteps), ncol=1), # Matrix of feeding success parameters per round of IRS and per species
ms_theta = matrix(-1.232, nrow=length(sprayingtimesteps), ncol=1), # Matrix of deterrence parameters per round of IRS and per species
ms_gamma = matrix(-0.009, nrow=length(sprayingtimesteps), ncol=1) # Matrix of deterrence parameters per round of IRS and per species
)
output <- run_simulation(timesteps = sim_length, parameters = sprayingparams)