Fitting odin models with monty
A pragmatic introduction
A script containing the code featured in these slides can be downloaded here.
Previously, on “Introduction to odin”
- We created some simple compartmental models
- We ran these and observed trajectories over time
- We saw that stochastic models produce a family of trajectories
The data
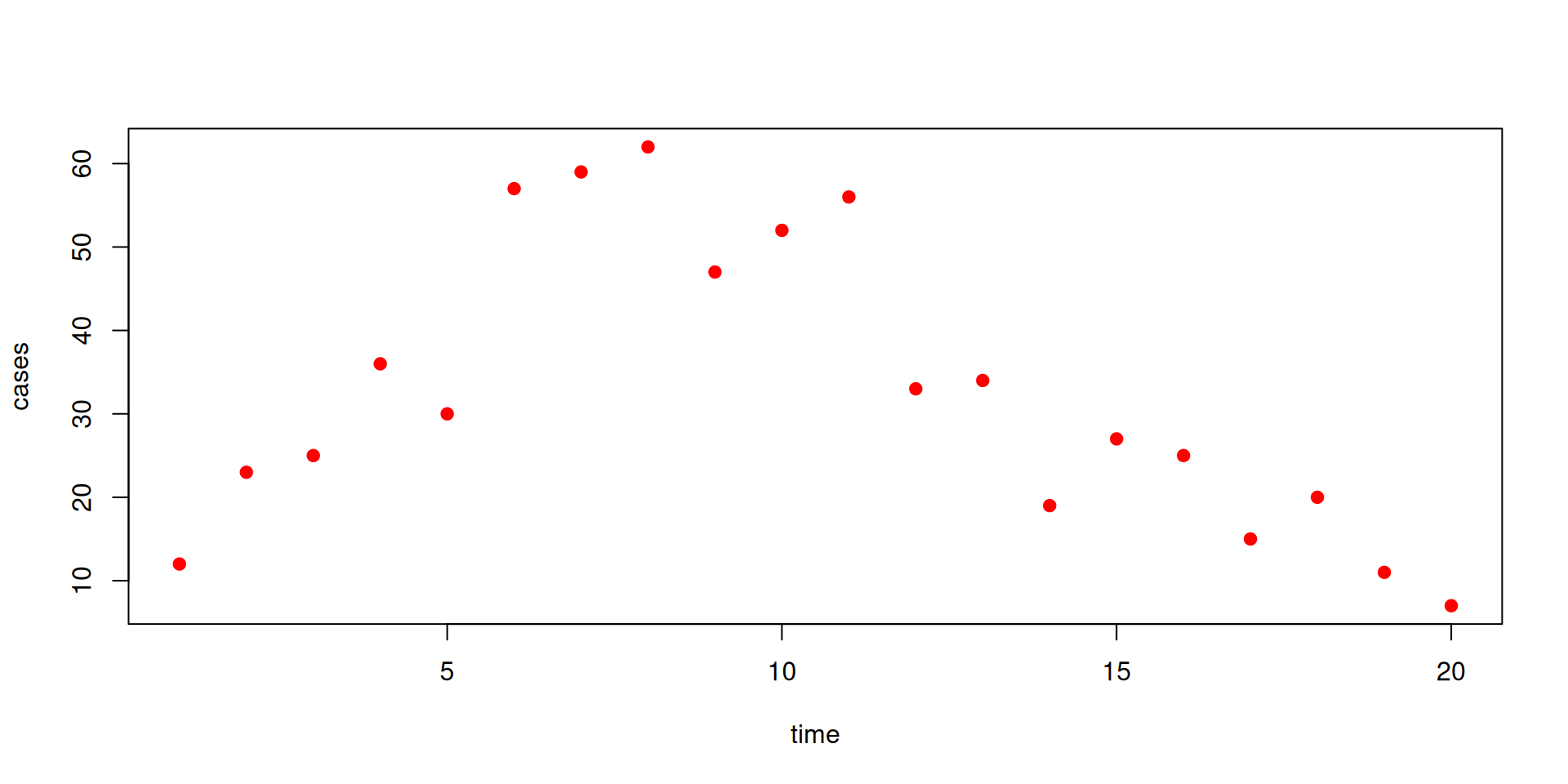
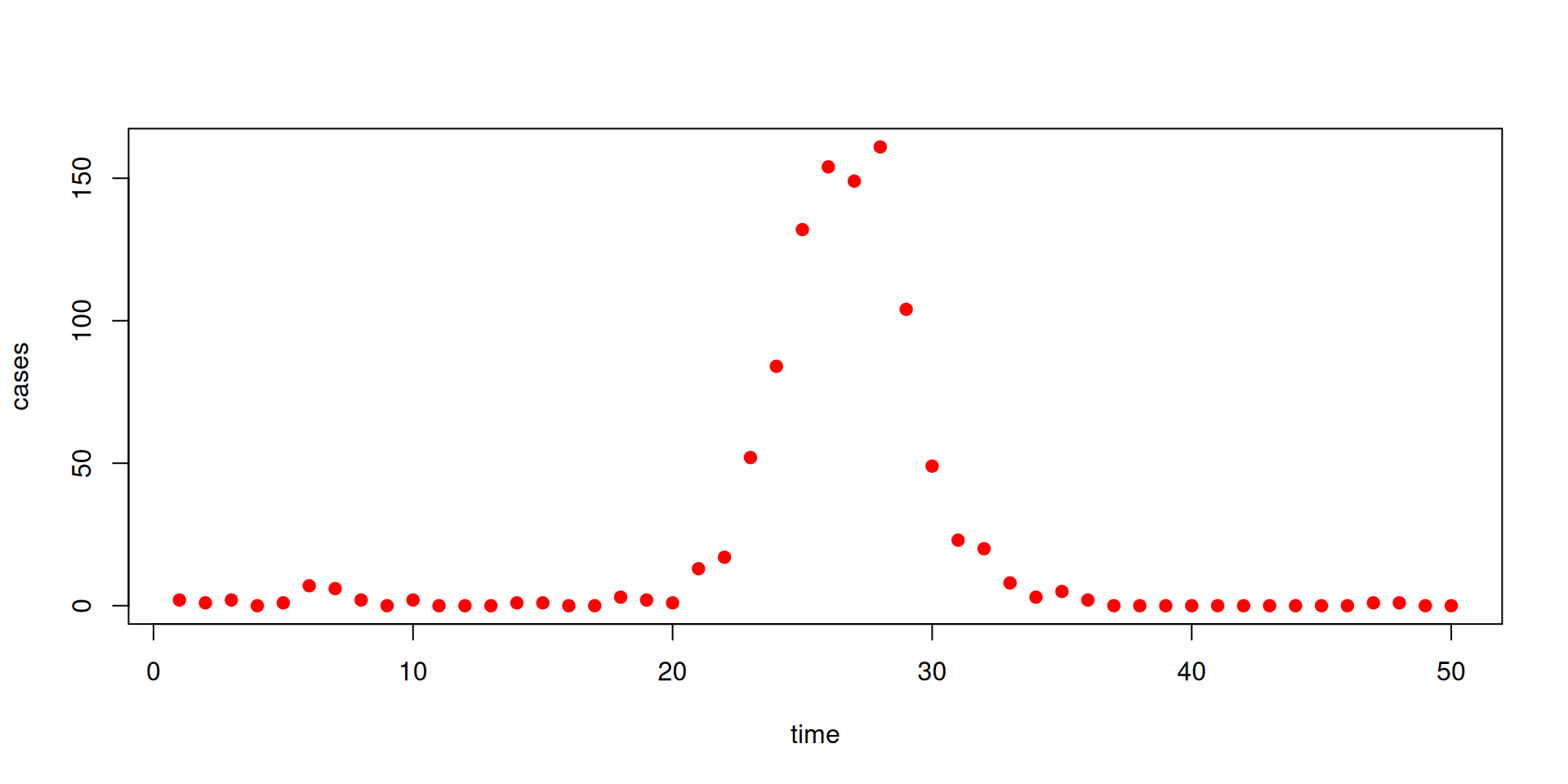
We have some data on the daily incidence of cases (the data can be downloaded here)
The data
Our model
We’ll fit these data to a model - estimating the unknown parameters beta and gamma
sir <- odin({
update(S) <- S - n_SI
update(I) <- I + n_SI - n_IR
update(R) <- R + n_IR
update(incidence) <- incidence + n_SI
p_SI <- 1 - exp(-beta * I / N * dt)
p_IR <- 1 - exp(-gamma * dt)
n_SI <- Binomial(S, p_SI)
n_IR <- Binomial(I, p_IR)
initial(S) <- N - I0
initial(I) <- I0
initial(R) <- 0
initial(incidence, zero_every = 1) <- 0
N <- parameter(1000)
I0 <- parameter(10)
beta <- parameter(0.2)
gamma <- parameter(0.1)
})Adding likelihood to the model
sir <- odin({
update(S) <- S - n_SI
update(I) <- I + n_SI - n_IR
update(R) <- R + n_IR
update(incidence) <- incidence + n_SI
initial(S) <- N - I0
initial(I) <- I0
initial(R) <- 0
initial(incidence, zero_every = 1) <- 0
p_SI <- 1 - exp(-beta * I / N * dt)
p_IR <- 1 - exp(-gamma * dt)
n_SI <- Binomial(S, p_SI)
n_IR <- Binomial(I, p_IR)
N <- parameter(1000)
I0 <- parameter(10)
beta <- parameter(0.2)
gamma <- parameter(0.1)
cases <- data()
cases ~ Poisson(incidence)
})Adding likelihood to the model
We have added to the model:
We are declaring that
casesis a data variable- it relates to
incidencein the model via a Poisson distribution
Note the two types of equations involving distributions:
~indicates a likelihood relationship<-is for assignments (that can involve random draws)
Beyond that syntax for distributions is the same.
In the book: Data
Adding likelihood to the model
You can use data variables on the RHS as well
and parameters
The distributions supported for likelihoods are the same as those supported for random draws - a full list with their names and parameterisations can be found here.
In the book: Data
State space models
- A state space model (SSM) is a mathematical framework for modelling a dynamical system.
- It is built around two processes:
- state equations that describes the evolution of some latent variables (also referred as “hidden” states) over time
- observation equations that relates the observations to the latent variables.
Can you be more precise?
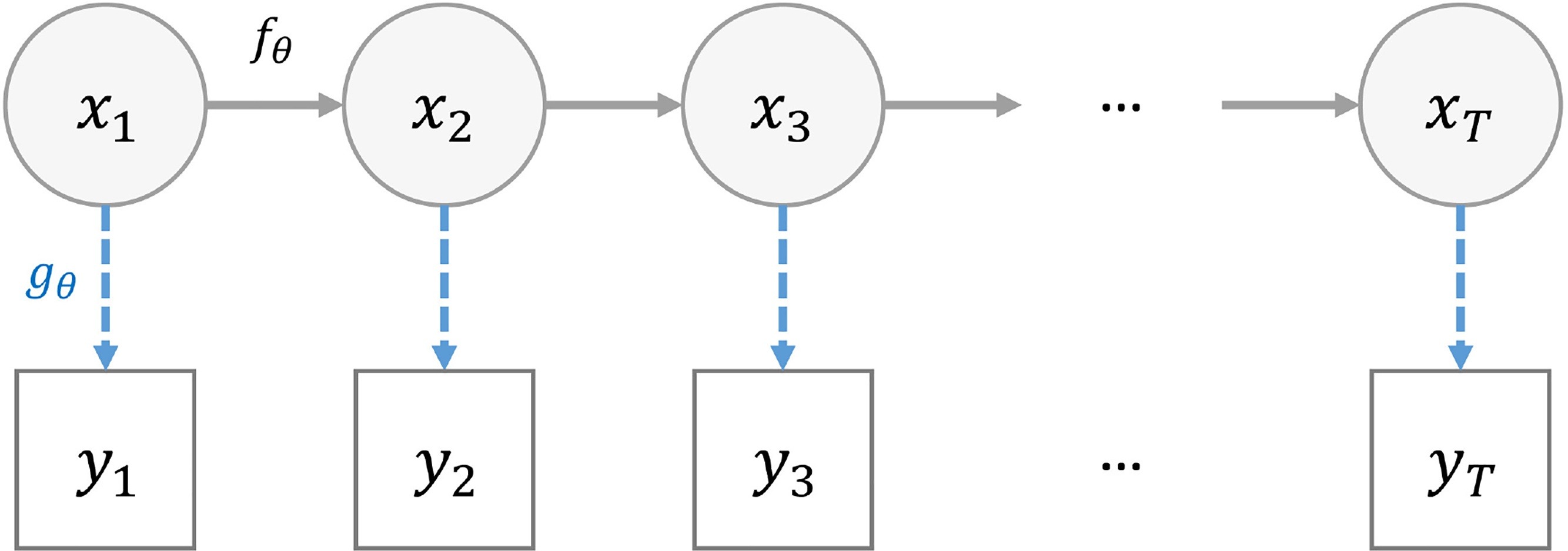
- \(x_{t, 1 \leq t \leq T}\) the hidden states of the system
- \(y_{t, 1 \leq t \leq T}\) the observations
- \(f_{\theta}\) the state transition function
- \(g_{\theta}\) the observation (likelihood) function
- \(t\) is often time
- \(\theta\) defines the model
Two common problems

- Two common needs
- “Filtering” i.e. estimate the hidden states \(x_{t}\) from the observations \(y_t\)
- “Inference” i.e. estimate the \(\theta\)’s compatible with the observations \(y_{t}\)
Sequential Monte Carlo (SMC)
AKA, the (bootstrap) particle filter
Assuming a given \(\theta\), at each time step \(t\) it
- generates \(X_{t+1}^N\) by using \(f_{\theta}(X_{t+1}^N|X_{t}^N)\) (the \(N\) particles)
- calculates weights for the newly generated states based on \(g_{\theta}(Y_{t+1}|X_{t+1})\)
- resamples the states to keep only the good ones
Allows to efficiently explore the state space by progressively integrating the data points
Produces a Monte Carlo approximation of \(p(Y_{1:T}|\theta)\) the marginal likelihood
Calculating likelihood: particle filtering
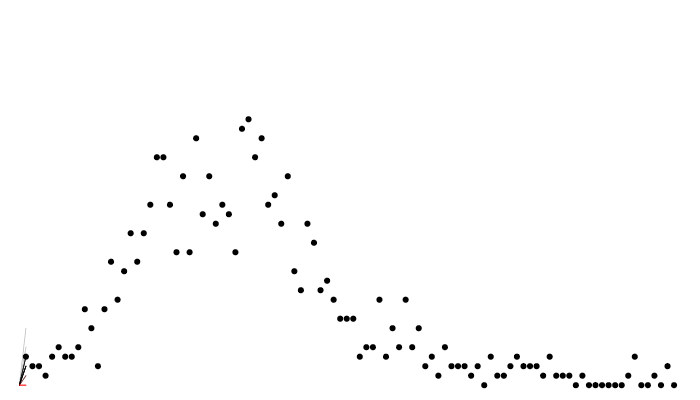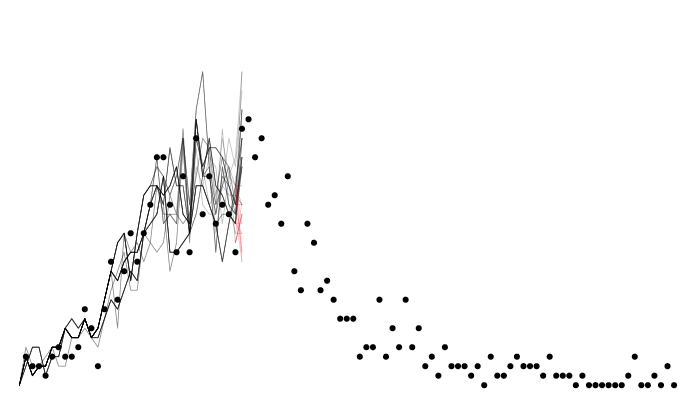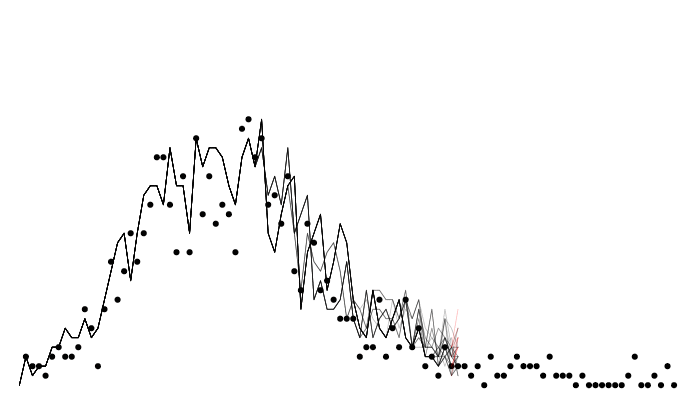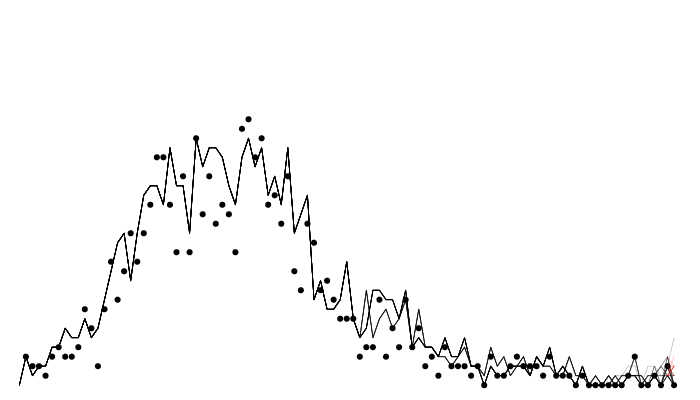
Calculating likelihood
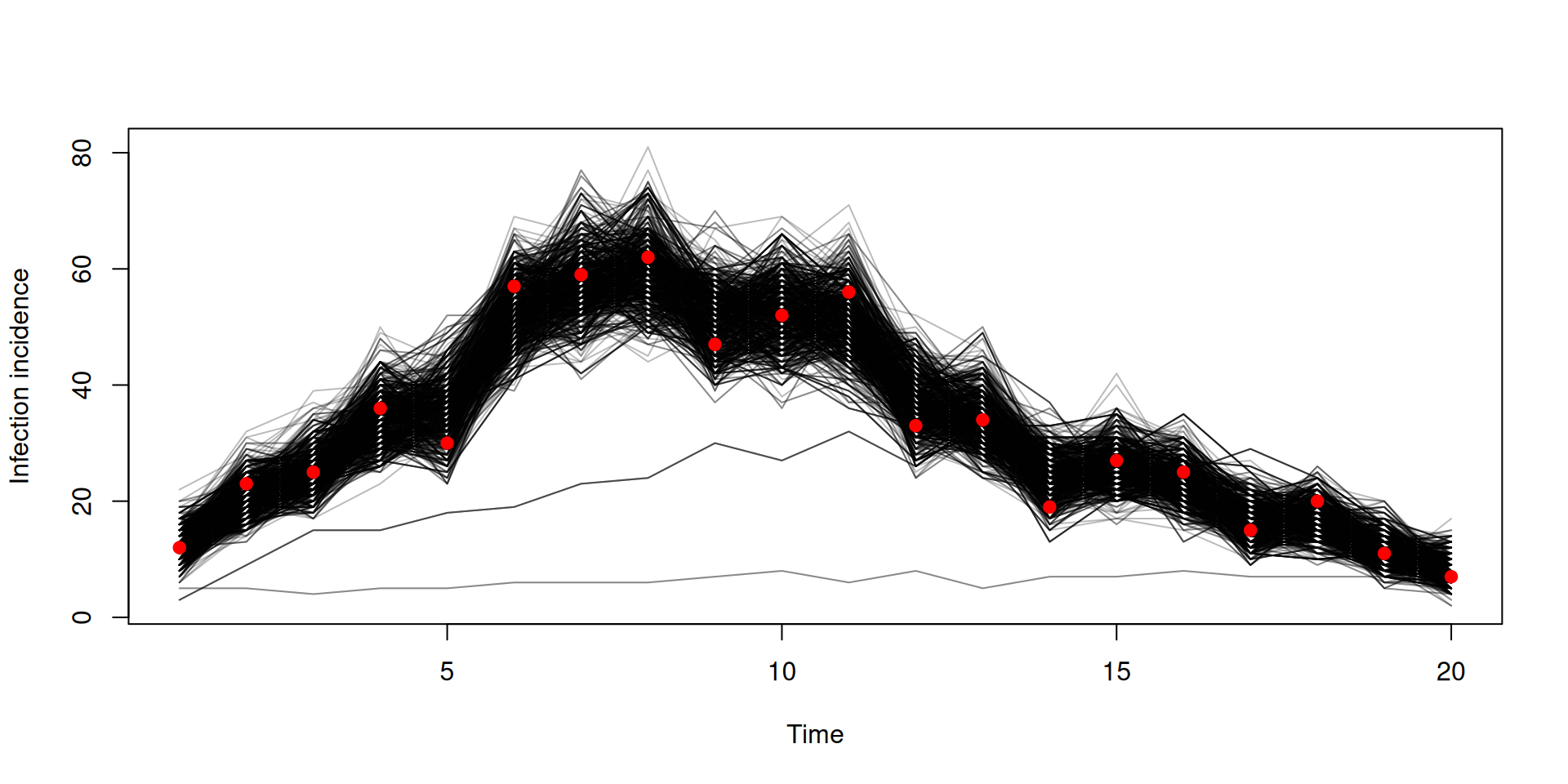
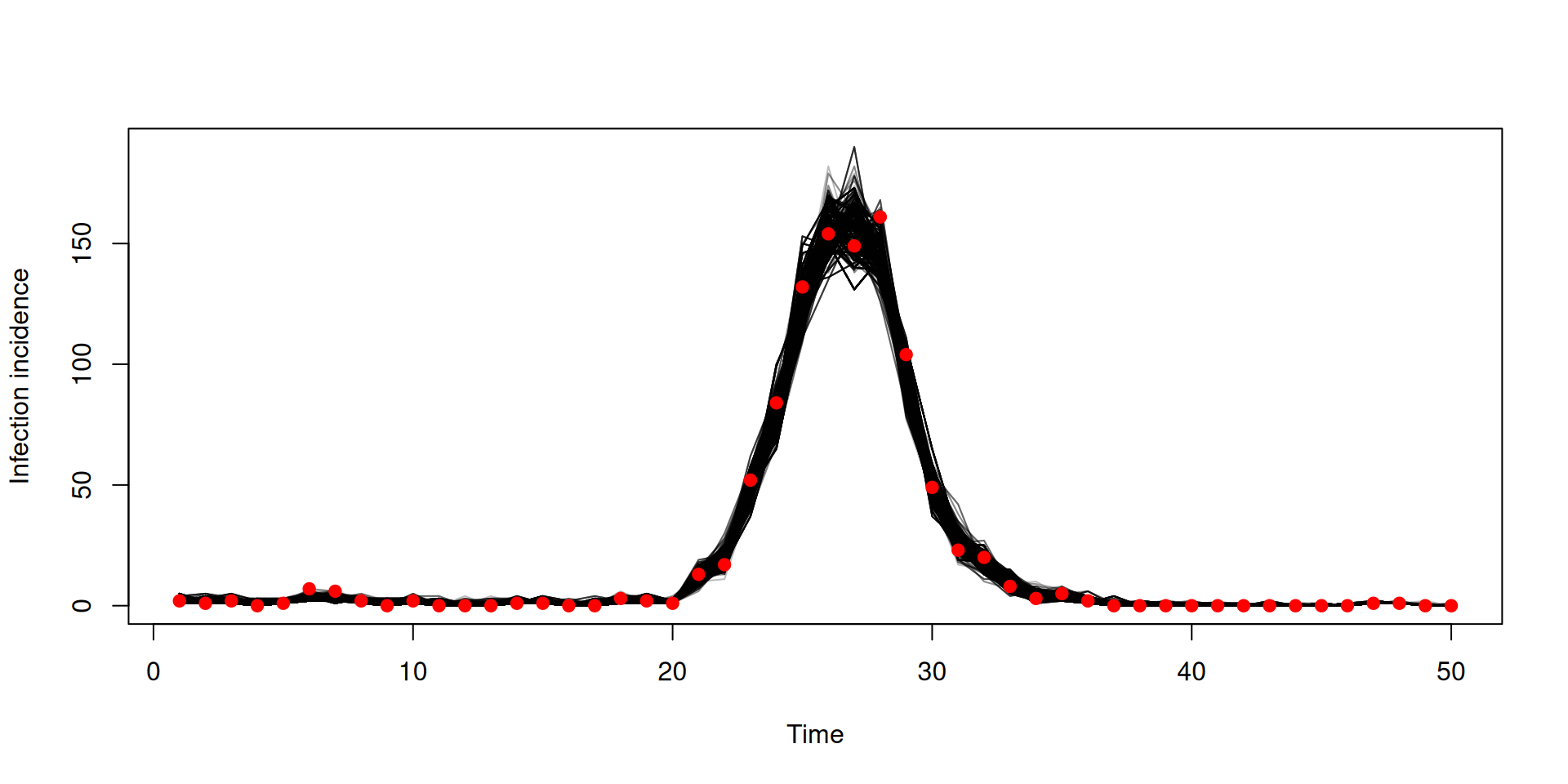
Filtered trajectories
dust_likelihood_run(filter, list(beta = 0.4, gamma = 0.2),
save_trajectories = TRUE)
y <- dust_likelihood_last_trajectories(filter)
y <- dust_unpack_state(filter, y)
matplot(data$time, t(y$incidence), type = "l", col = "#00000044", lty = 1,
xlab = "Time", ylab = "Incidence")
points(data, pch = 19, col = "red")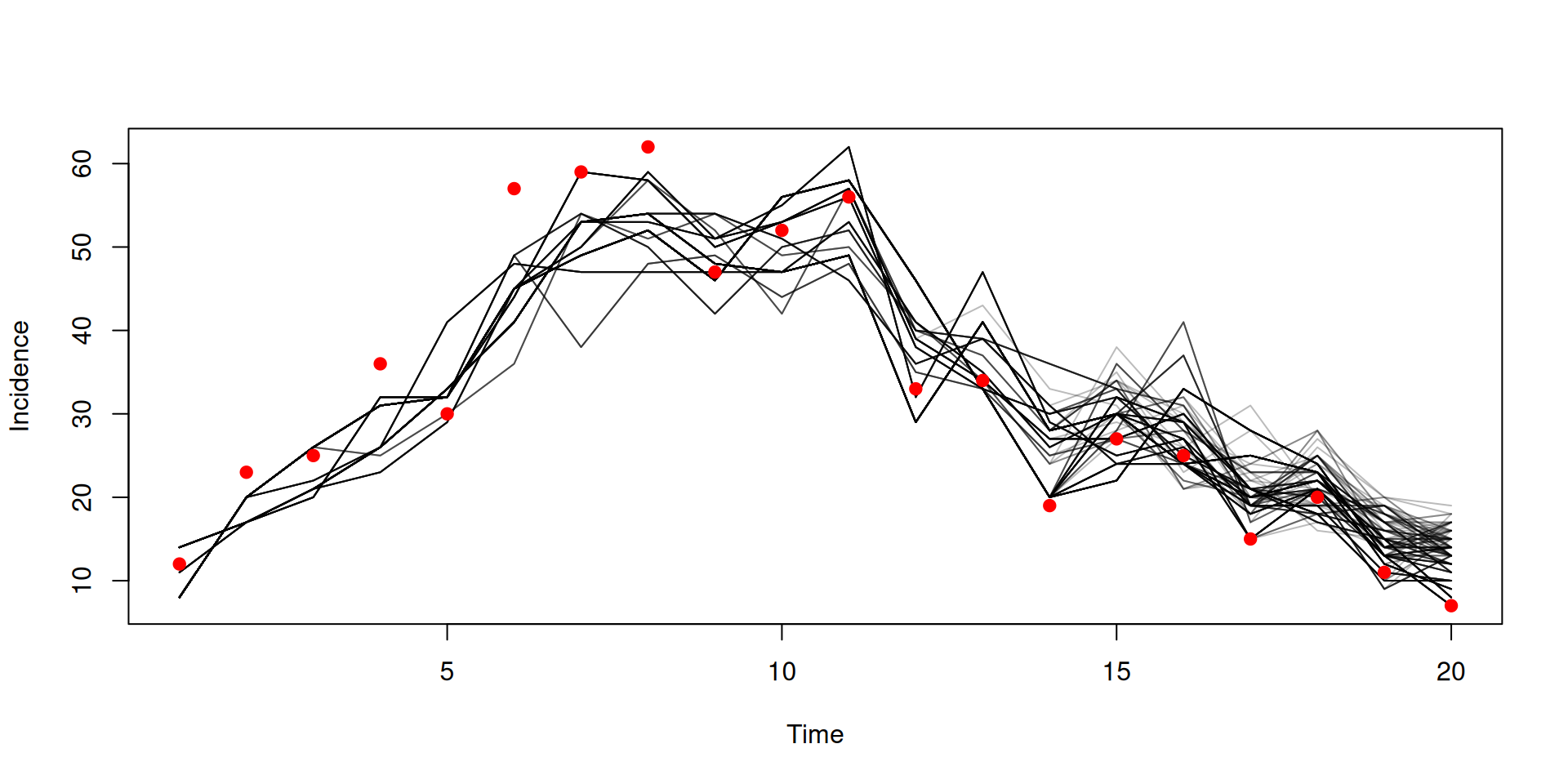
Particle MCMC (PMCMC)
- PMCMC is an algorithm which performs “filtering” and “inference”
- A Markov Chain Monte Carlo (MCMC) method for estimating target distributions
- MCMC explores the parameter space by moving randomly making jumps from one value to the next
- In PMCMC, the marginal likelihood is estimated using a particle filter
- One particle trajectory is selected randomly at the end of the particle filter
Particle MCMC
So we have a marginal likelihood estimator from our particle filter
How do we sample from beta and gamma?
We need:
- to tidy up our parameters
- to create a prior
- to create a posterior
- to create a sampler
“Parameters”
- Our filter takes a list of
betaandgamma,pars- it could take all sorts of other things, not all of which are to be estimated
- some of the inputs might be vectors or matrices
- Our MCMC takes an unstructured vector \(\theta\)
- we propose a new \(\theta^*\) via some kernel, say a multivariate normal requiring a matrix of parameters corresponding to \(\theta\)
- we need a prior over \(\theta\), but not necessarily every element of
pars
- Smoothing this over is a massive nuisance
- some way of mapping from \(\theta\) to
pars(and back again)
- some way of mapping from \(\theta\) to
Parameter packers
Our solution, “packers”
packer <- monty_packer(c("beta", "gamma"))
packer
#>
#> ── <monty_packer> ──────────────────────────────────────────────────────────────
#> ℹ Packing 2 parameters: 'beta' and 'gamma'
#> ℹ Use '$pack()' to convert from a list to a vector
#> ℹ Use '$unpack()' to convert from a vector to a list
#> ℹ See `?monty_packer()` for more informationParameter packers
We can transform from \(\theta\) to a named list:
and back the other way:
Parameter packers
You can bind fixed parameters by passing them in as a named list as the (optional) fixed argument
Parameter packers
Cope with vector- (or array-) parameters in \(\theta\) by declaring their dimensions as a named list passed in as the array argument. Fixed array parameters are just included in fixed.
packer <- monty_packer(c("beta", "gamma"),
array = list(alpha = 3, delta = c(2, 2)),
fixed = list(eta = c(1, 3, 5)))
packer
#>
#> ── <monty_packer> ──────────────────────────────────────────────────────────────
#> ℹ Packing 9 parameters: 'beta', 'gamma', 'alpha[1]', 'alpha[2]', 'alpha[3]', 'delta[1,1]', 'delta[2,1]', 'delta[1,2]', and 'delta[2,2]'
#> ℹ Use '$pack()' to convert from a list to a vector
#> ℹ Use '$unpack()' to convert from a vector to a list
#> ℹ See `?monty_packer()` for more information
packer$unpack(c(0.2, 0.1, 0.31, 0.32, 0.33, 0.15, 0.16, 0.17, 0.18))
#> $beta
#> [1] 0.2
#>
#> $gamma
#> [1] 0.1
#>
#> $alpha
#> [1] 0.31 0.32 0.33
#>
#> $delta
#> [,1] [,2]
#> [1,] 0.15 0.17
#> [2,] 0.16 0.18
#>
#> $eta
#> [1] 1 3 5The monty DSL
Working in a Bayesian framework, we will need to construct a prior distribution model for our parameters.
Introducing the monty DSL, similar to odin’s:
The distributions supported in the monty DSL are the same as those in odin2 - a full list with their names and parameterisations can be found here.
In the book: The monty DSL
The monty DSL
The monty DSL creates a “monty model”
In the book: The monty DSL
The monty DSL
You can see the parameter names of the created model
We can evaluate the (log) density for a given parameter vector (the order of parameters matches the above)
compare with computing this density manually:
In the book: The monty DSL
The monty DSL
We can direct sample from the model using a monty_rng object
and evaluate the gradient for a given parameter vector (via automatic differentiation)
Which can be used for Hamiltonian Monte Carlo (HMC) - more support for this in future!
In the book: The monty DSL
The monty DSL
In the monty DSL you can have the distribution of one parameter depend upon another
m <- monty_dsl({
a ~ Normal(0, 1)
b ~ Normal(a, 1)
})
m
#>
#> ── <monty_model> ───────────────────────────────────────────────────────────────
#> ℹ Model has 2 parameters: 'a' and 'b'
#> ℹ This model:
#> • can compute gradients
#> • can be directly sampled from
#> • accepts multiple parameters
#> ℹ See `?monty_model()` for more informationIn the book: The monty DSL
The monty DSL
Order matters in the monty DSL - you cannot have the distribution of a parameter depend upon one defined later, so switching the order in our previous model results in an error
In the book: The monty DSL
The monty DSL
You can use assignments to help make your code more understandable
these can also be used for intermediate calculations
In the book: The monty DSL
The monty DSL
You can softcode values by passing them in as a list via fixed, e.g.
can be implemented as
Note that the fixed values cannot be modified after the model object has been created.
In the book: The monty DSL
The monty DSL
The monty DSL supports arrays with a similar syntax to odin.
fixed = list(gamma_mean = c(0.3, 0.25, 0.2))
m <- monty_dsl({
alpha ~ Exponential(mean = 0.5)
beta[, ] ~ Normal(0, 1)
gamma[] ~ Exponential(mean = gamma_mean[i])
dim(beta) <- c(2, 2)
dim(gamma, gamma_mean) <- 3
}, fixed = fixed, gradient = FALSE)
m
#>
#> ── <monty_model> ───────────────────────────────────────────────────────────────
#> ℹ Model has 8 parameters: 'alpha', 'beta[1,1]', 'beta[2,1]', 'beta[1,2]', 'beta[2,2]', 'gamma[1]', 'gamma[2]', and 'gamma[3]'
#> ℹ This model:
#> • can be directly sampled from
#> • accepts multiple parameters
#> ℹ See `?monty_model()` for more informationIn the book: The monty DSL
The monty DSL
- All arrays need a
dim()equation - No use of
i,jetc on LHS - indexing on the LHS is implicit - Support for up to 8 dimensions, with index variables
i,j,k,l,i5,i6,i7,i8
essentially generates loops
In the book: The monty DSL
The monty DSL
For flexibility dimensions can be softcoded and passed in via fixed
fixed = list(gamma_mean = c(0.3, 0.25, 0.2),
n_beta_1 = 2, n_beta_2 = 2, n_gamma = 3)
m <- monty_dsl({
alpha ~ Exponential(mean = 0.5)
beta[, ] ~ Normal(0, 1)
gamma[] ~ Exponential(mean = gamma_mean[i])
dim(beta) <- c(n_beta_1, n_beta_2)
dim(gamma, gamma_mean) <- n_gamma
}, fixed = fixed, gradient = FALSE)
m
#>
#> ── <monty_model> ───────────────────────────────────────────────────────────────
#> ℹ Model has 8 parameters: 'alpha', 'beta[1,1]', 'beta[2,1]', 'beta[1,2]', 'beta[2,2]', 'gamma[1]', 'gamma[2]', and 'gamma[3]'
#> ℹ This model:
#> • can be directly sampled from
#> • accepts multiple parameters
#> ℹ See `?monty_model()` for more informationIn the book: The monty DSL
The monty DSL
You can define arrays with multiline equations
but collectively the equations need to define all elements of the array, and each element can only be defined once.
In the book: The monty DSL
The monty DSL
Elements of an array can depend upon other elements of the same array that are defined earlier.
We can see that in our previous example
which becomes clearer when we think how the loop will be generated
In the book: The monty DSL
From a dust filter to a monty model
Combine a filter and a packer
packer <- monty_packer(c("beta", "gamma"))
likelihood <- dust_likelihood_monty(filter, packer)
likelihood
#>
#> ── <monty_model> ───────────────────────────────────────────────────────────────
#> ℹ Model has 2 parameters: 'beta' and 'gamma'
#> ℹ This model:
#> • is stochastic
#> ℹ See `?monty_model()` for more informationIn the book: Inference
Posterior from likelihood and prior
Combine a likelihood and a prior to make a posterior
\[ \underbrace{\Pr(\theta | \mathrm{data})}_{\mathrm{posterior}} \propto \underbrace{\Pr(\mathrm{data} | \theta)}_\mathrm{likelihood} \times \underbrace{P(\theta)}_{\mathrm{prior}} \]
(remember that addition is multiplication on a log scale)
Create a sampler
A diagonal variance-covariance matrix (uncorrelated parameters)
Use this to create a “random walk” sampler:
In the book: Samplers and inference
Let’s sample!
We will sample 4 chains, each with 1000 steps
samples <- monty_sample(posterior, sampler, 1000, n_chains = 3)
samples
#>
#> ── <monty_samples: 2 parameters x 1000 samples x 3 chains> ─────────────────────
#> ℹ Parameters: 'beta' and 'gamma'
#> ℹ Conversion to other types is possible:
#> → ! posterior::as_draws_array() [package installed, but not loaded]
#> → ! posterior::as_draws_df() [package installed, but not loaded]
#> → ! coda::as.mcmc.list() [package installed, but not loaded]
#> ℹ See `?monty_sample()` and `vignette("samples")` for more informationIn the book: Inference
The result: diagnostics
Diagnostics can be used from the posterior package
## Note: as_draws_df converts samples$pars, and drops anything else in samples
samples_df <- posterior::as_draws_df(samples)
posterior::summarise_draws(samples_df)
#> # A tibble: 2 × 10
#> variable mean median sd mad q5 q95 rhat ess_bulk ess_tail
#> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 beta 0.866 0.831 0.153 0.152 0.658 1.15 1.05 50.8 68.8
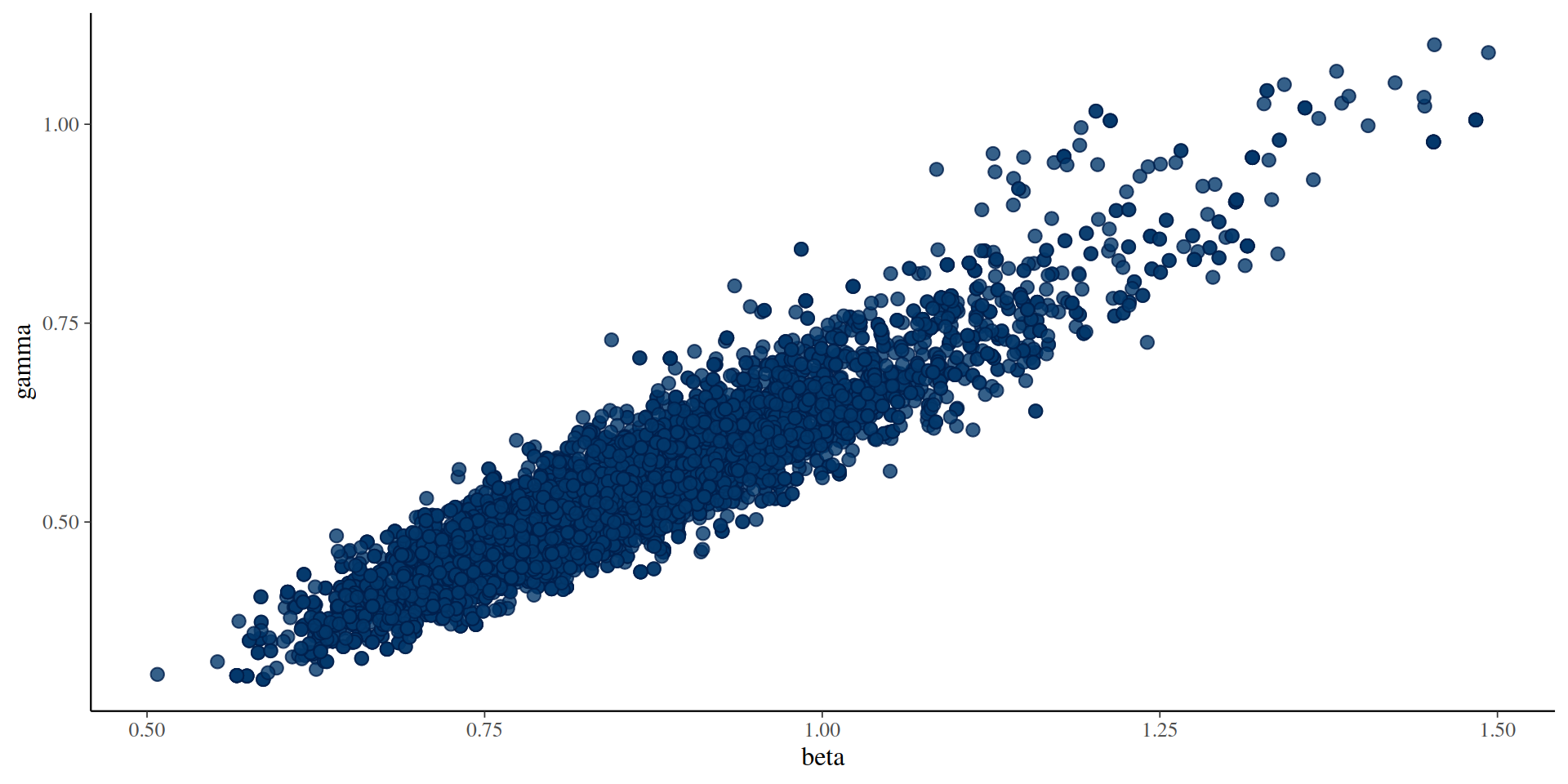
#> 2 gamma 0.549 0.545 0.119 0.115 0.381 0.728 1.04 50.5 77.1The results: parameters
You can use the posterior package in conjunction with bayesplot (and then also ggplot2)
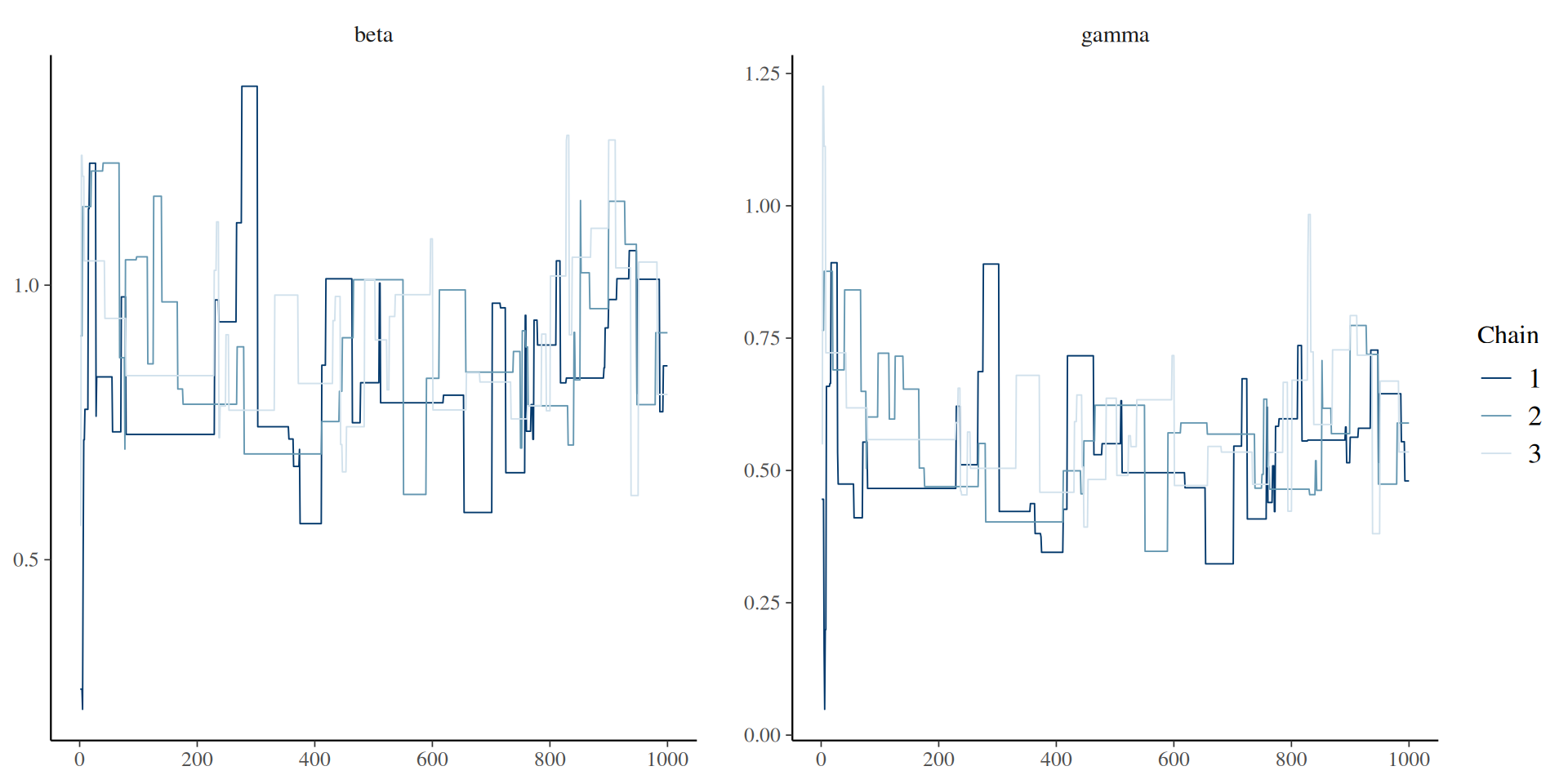
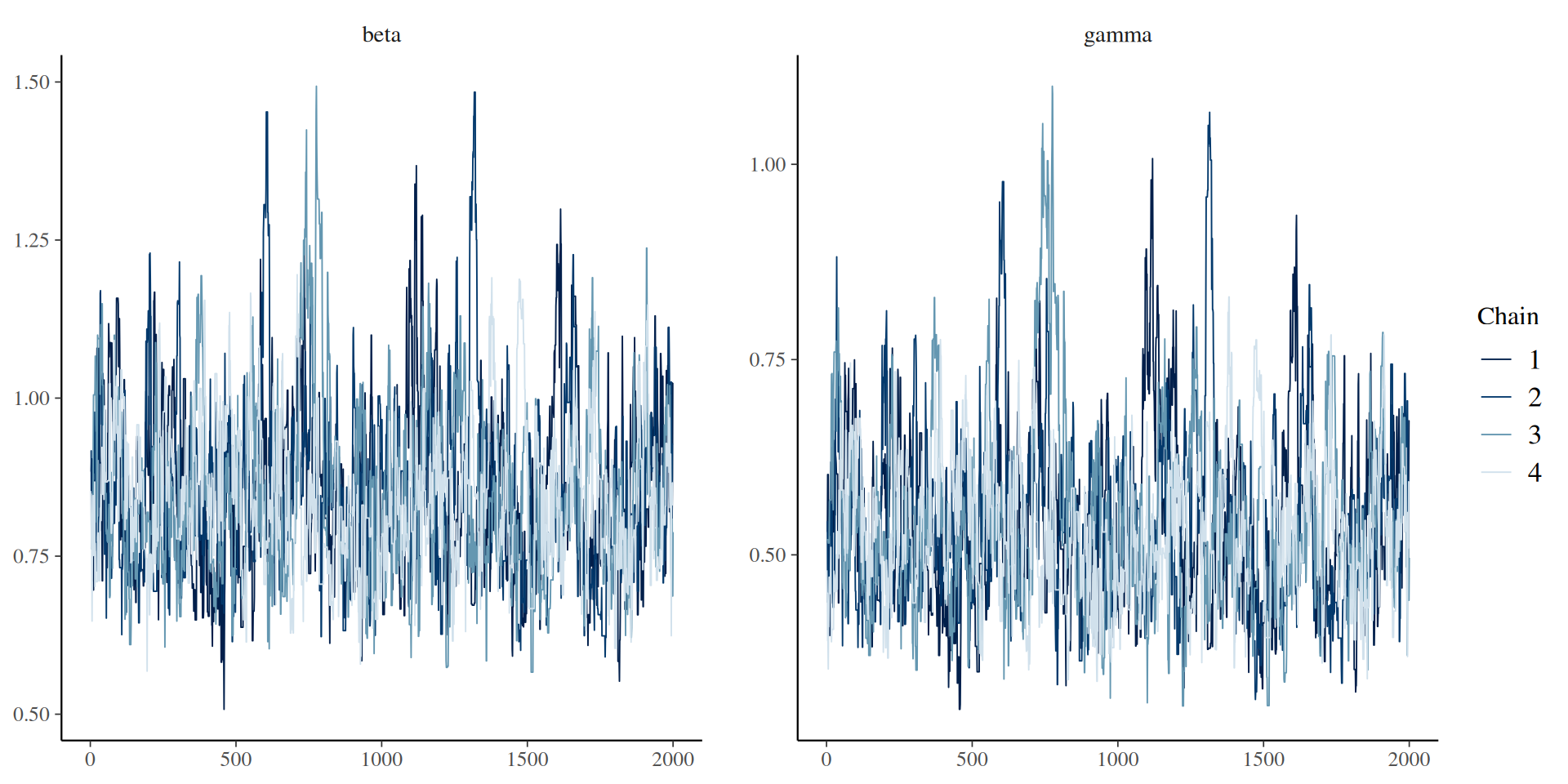
The result: traceplots
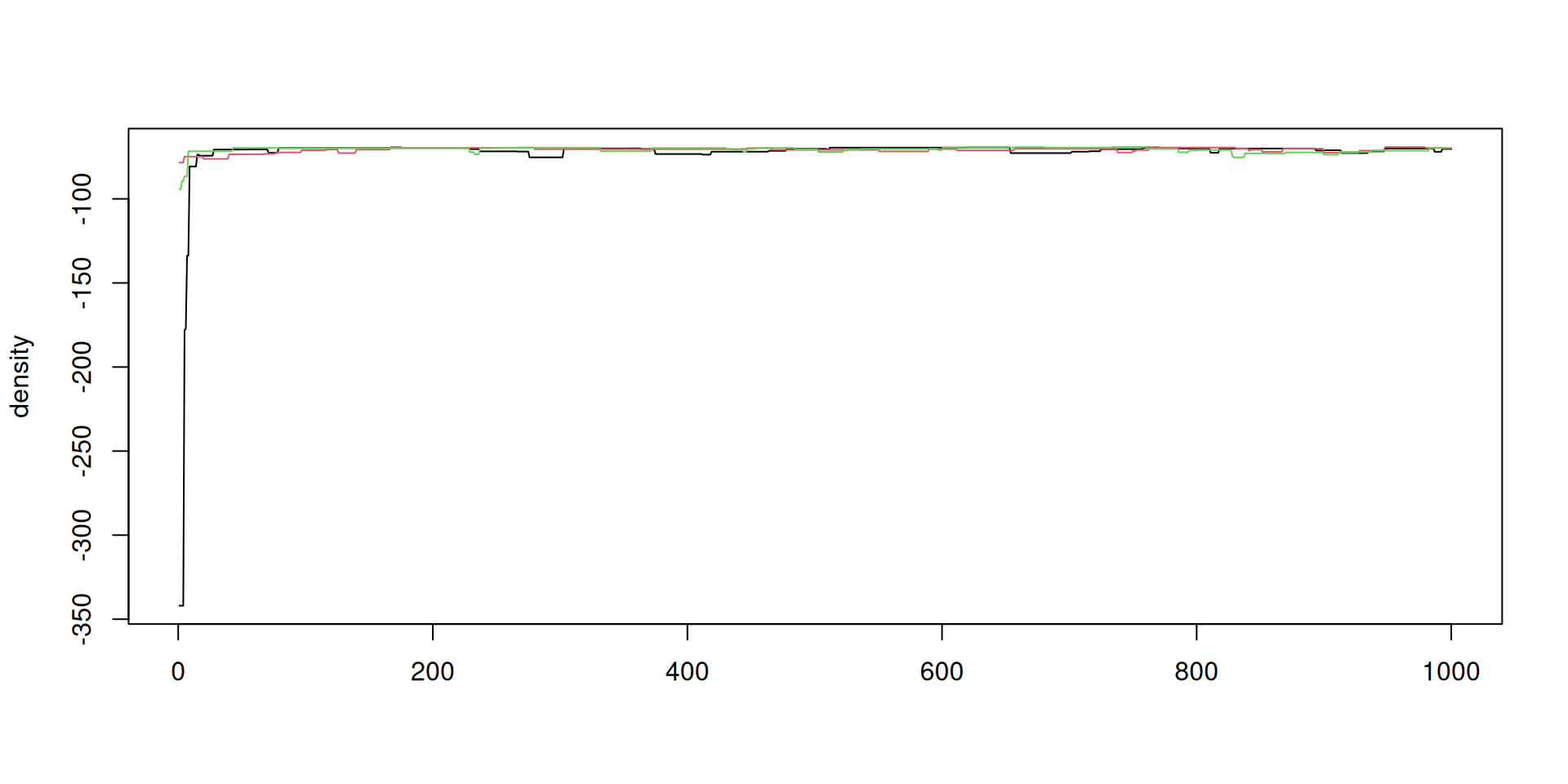
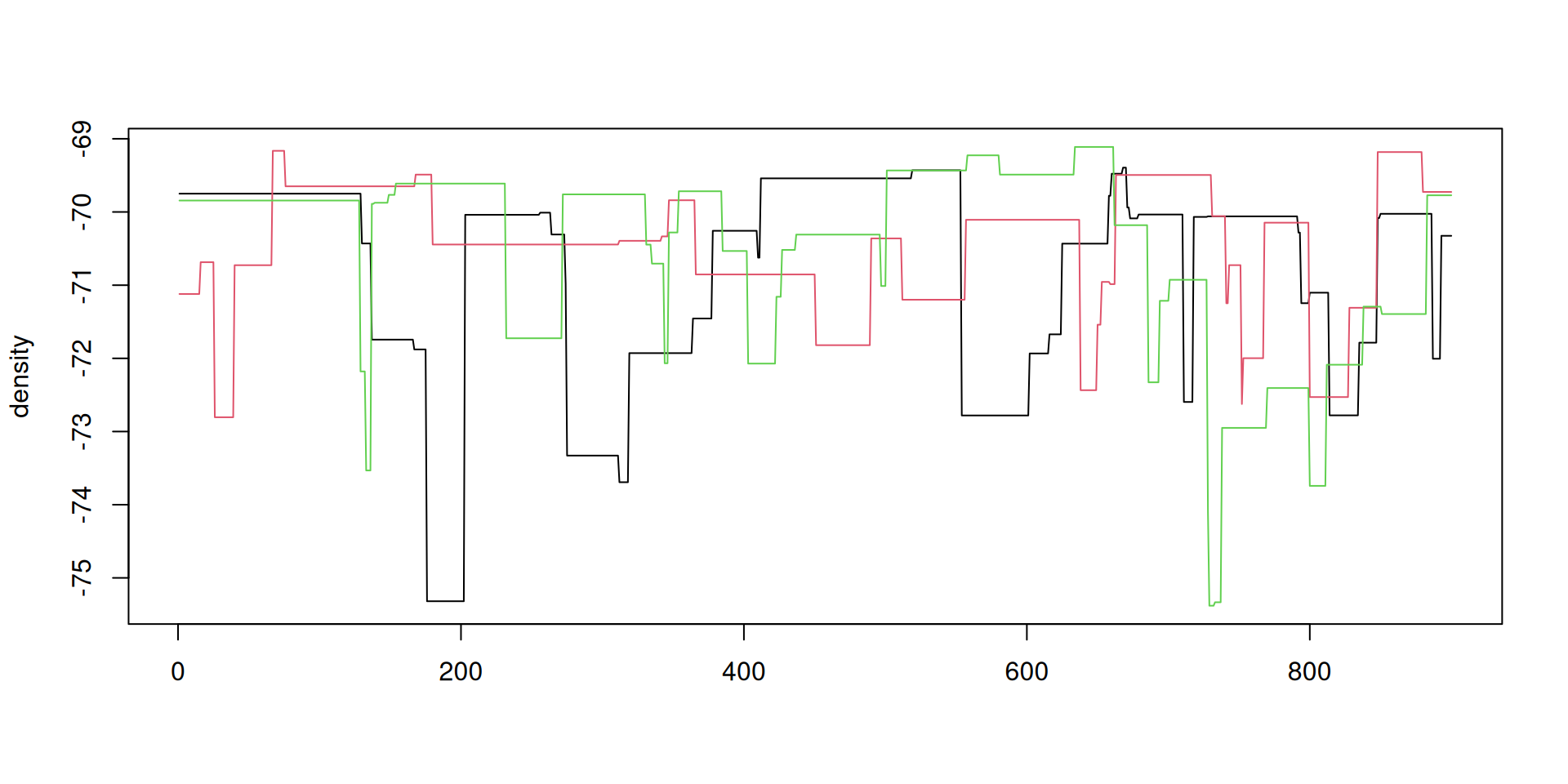
The result: density over time
The result: density over time
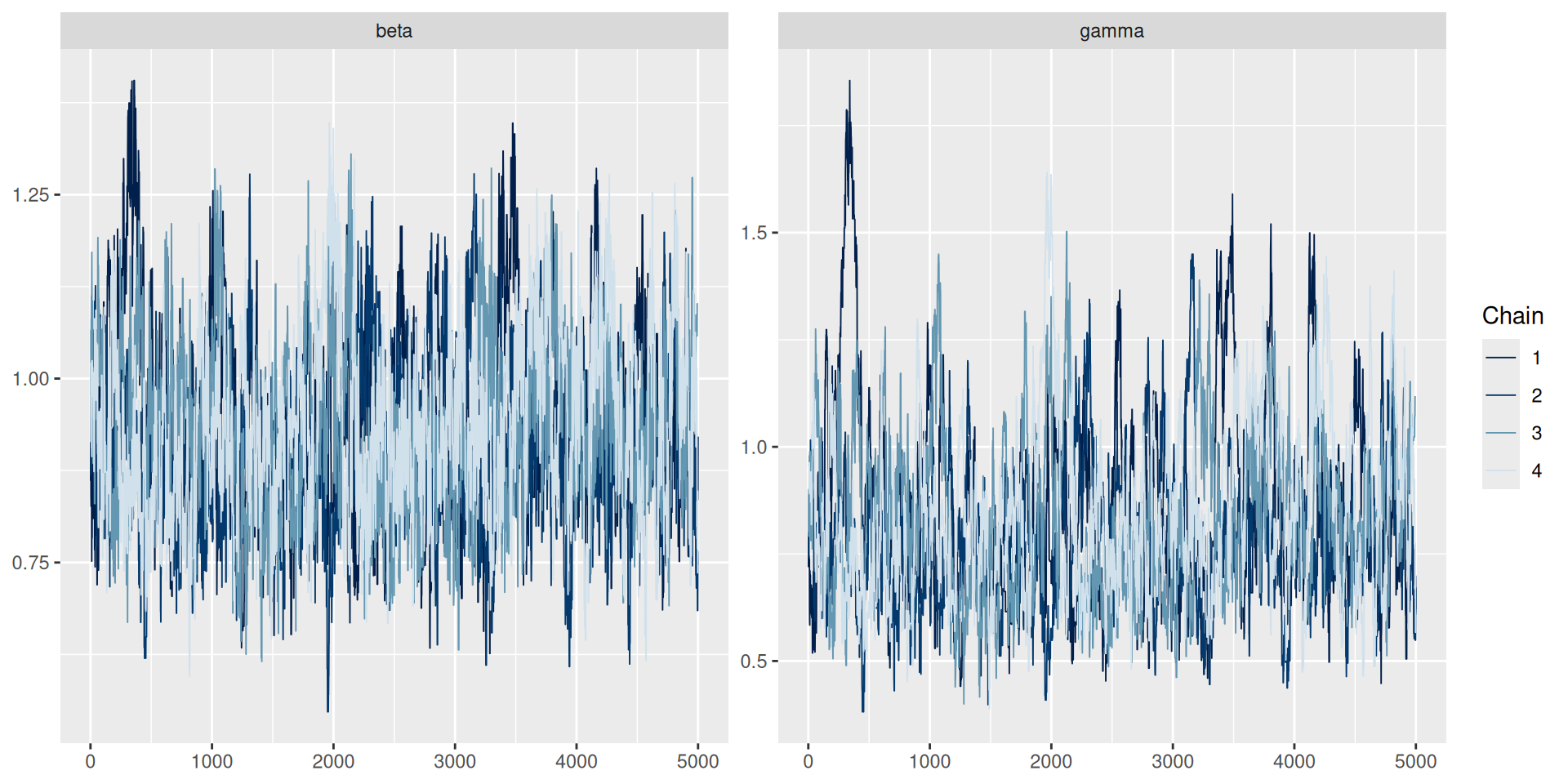
Better mixing
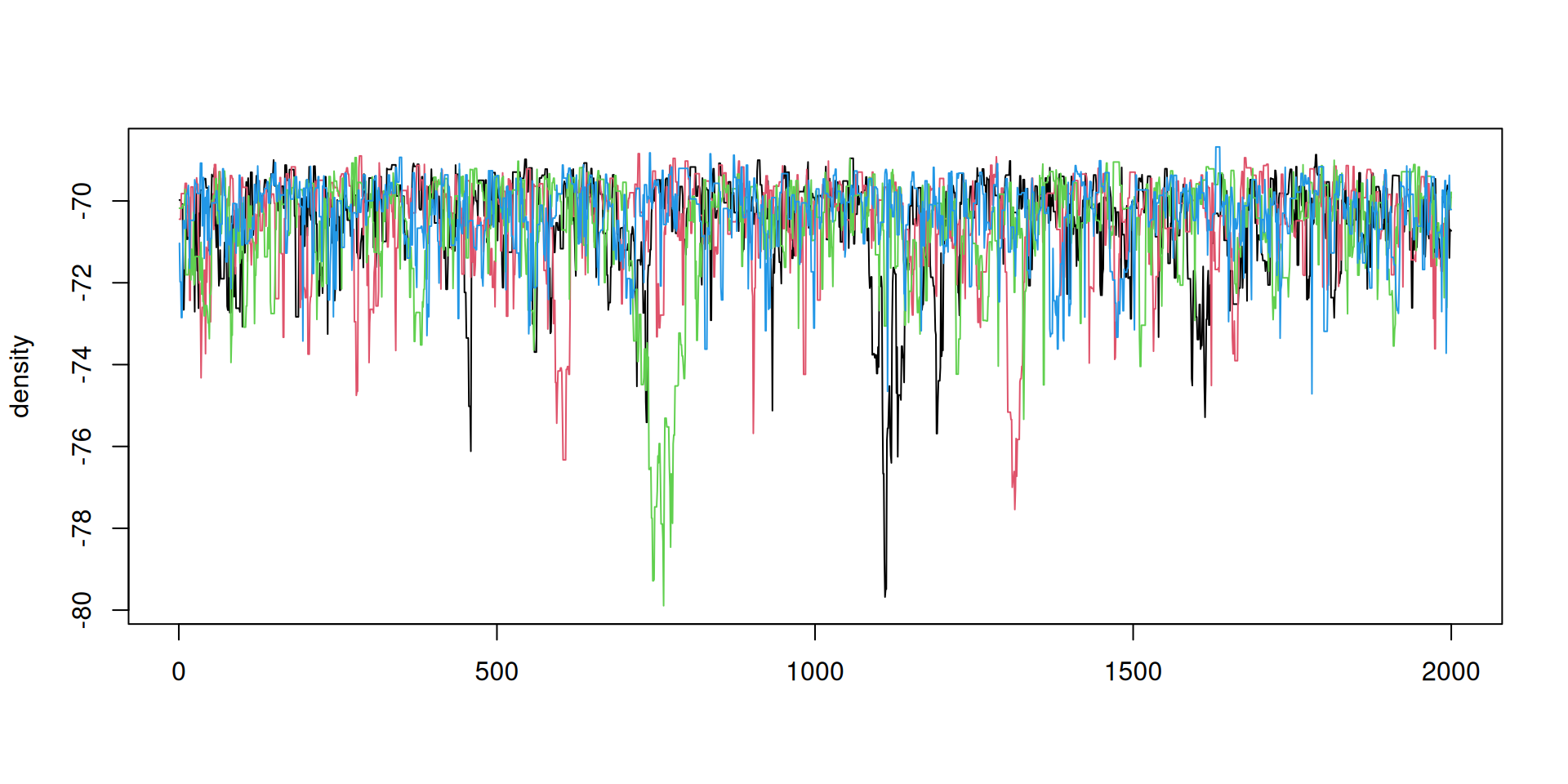
Better mixing: the results
samples_df <- posterior::as_draws_df(samples)
posterior::summarise_draws(samples_df)
#> # A tibble: 2 × 10
#> variable mean median sd mad q5 q95 rhat ess_bulk ess_tail
#> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 beta 0.851 0.835 0.131 0.120 0.667 1.09 1.01 338. 388.
#> 2 gamma 0.539 0.524 0.107 0.0944 0.393 0.737 1.00 301. 286.Better mixing: the results
Better mixing: the results
Parallelism
Two places to parallelise
- among particles in your filter
- between chains in the sample
e.g., 4 threads per filter x 2 workers = 8 total cores in use
In the book: Parallelisation
Configure the filter
Use the n_threads argument, here for 4 threads
requires that you have OpenMP.
In the book: Parallelisation
Configure a parallel runner
Use monty_runner_callr, here for 2 workers
Pass runner through to monty_sample:
In the book: Parallelisation
Run chains on different cluster nodes
Then run these chains in parallel on your cluster:
And retrieve the result
In the book: Parallelisation
Saving history
- Save your trajectories at every collected sample
- Save the final state at every sample (for onward simulation)
- Save snapshots at intermediate timepoints of the state at every sample (for counterfactuals)
We look at how to do onward simulation and counterfactuals later.
Trajectories
Trajectories
Trajectories
Trajectories are 4-dimensional
These can get very large quickly - there are two main ways to help reduce this:
- Saving only a subset of the states
- Thinning
Saving a subset of trajectories
You can save a subset via specifying a named vector
Thinning
After running
- Thinning while running faster and uses less memory
- After running is more flexible (e.g. can plot full chains of parameters between running and thinning)
Dealing with sticky chains
- Chains can get stuck when the variance of the marginal likelihood estimate is large
- The chain has obtained a high-end marginal likelihood estimate at one parameter set and needs another high-end estimate at another parameter set to accept
- You can increase the number of particles to reduce variance, but note this will take longer to run
- In
monty_sampler_random_walkyou can rerun the particle filter at the current parameter set with the aim of calculating a lower estimatererun_everysets how often you rerun (rerun_every = 1000means once every 1000 iterations)rerun_randomdefines whether this occurs randomly (TRUE) or at fixed intervals (FALSE)- using the rerun comes at cost of some bias in the results
- also some extra runtime cost (increasing the more frequently you rerun)
Deterministic models from stochastic
- Stochastic models written in odin, can be run deterministically
- Runs by taking the expectation of any random draws
- This gives two models for the price of one
- However it might not be suitable for all models
Fitting in deterministic mode
The key difference is to use dust_unfilter_create
Note as this is deterministic it produces the same likelihood every time
You should use the same setup if you have an ODE model (which is automatically deterministic)
Fitting in deterministic mode
Stochastic v deterministic comparison
y <- dust2::dust_unpack_state(filter, samples$observations$trajectories)
incidence <- array(y$incidence, c(20, 1000))
matplot(data$time, incidence, type = "l", lty = 1, col = "#00000044",
xlab = "Time", ylab = "Infection incidence", ylim = c(0, 75),
main = "Stochastic fit")
points(data, pch = 19, col = "red")
y <- dust2::dust_unpack_state(unfilter, samples_det$observations$trajectories)
incidence <- array(y$incidence, c(20, 1000))
matplot(data$time, incidence, type = "l", lty = 1, col = "#00000044",
xlab = "Time", ylab = "Infection incidence", ylim = c(0, 75),
main = "Deterministic fit")
points(data, pch = 19, col = "red")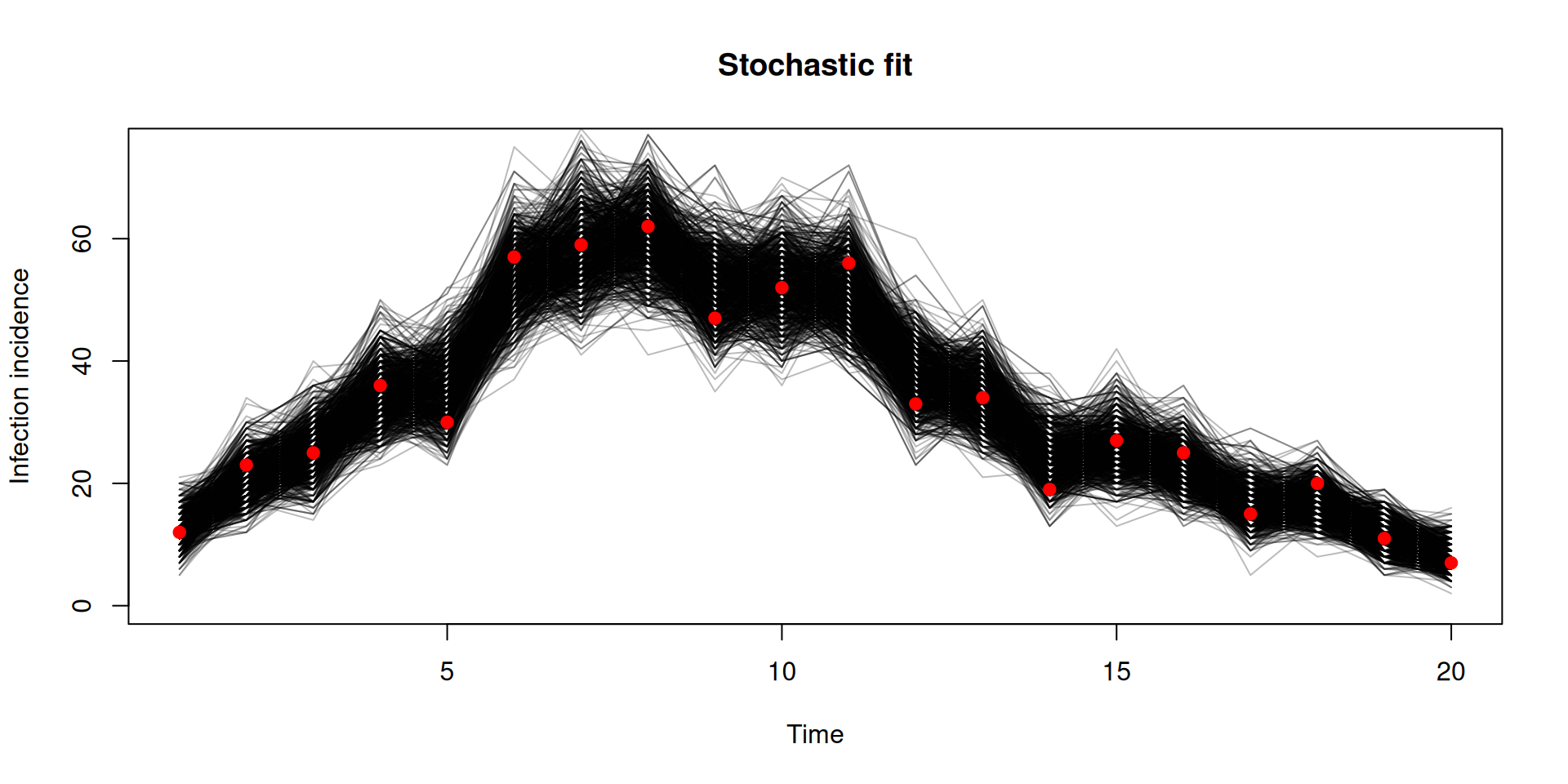
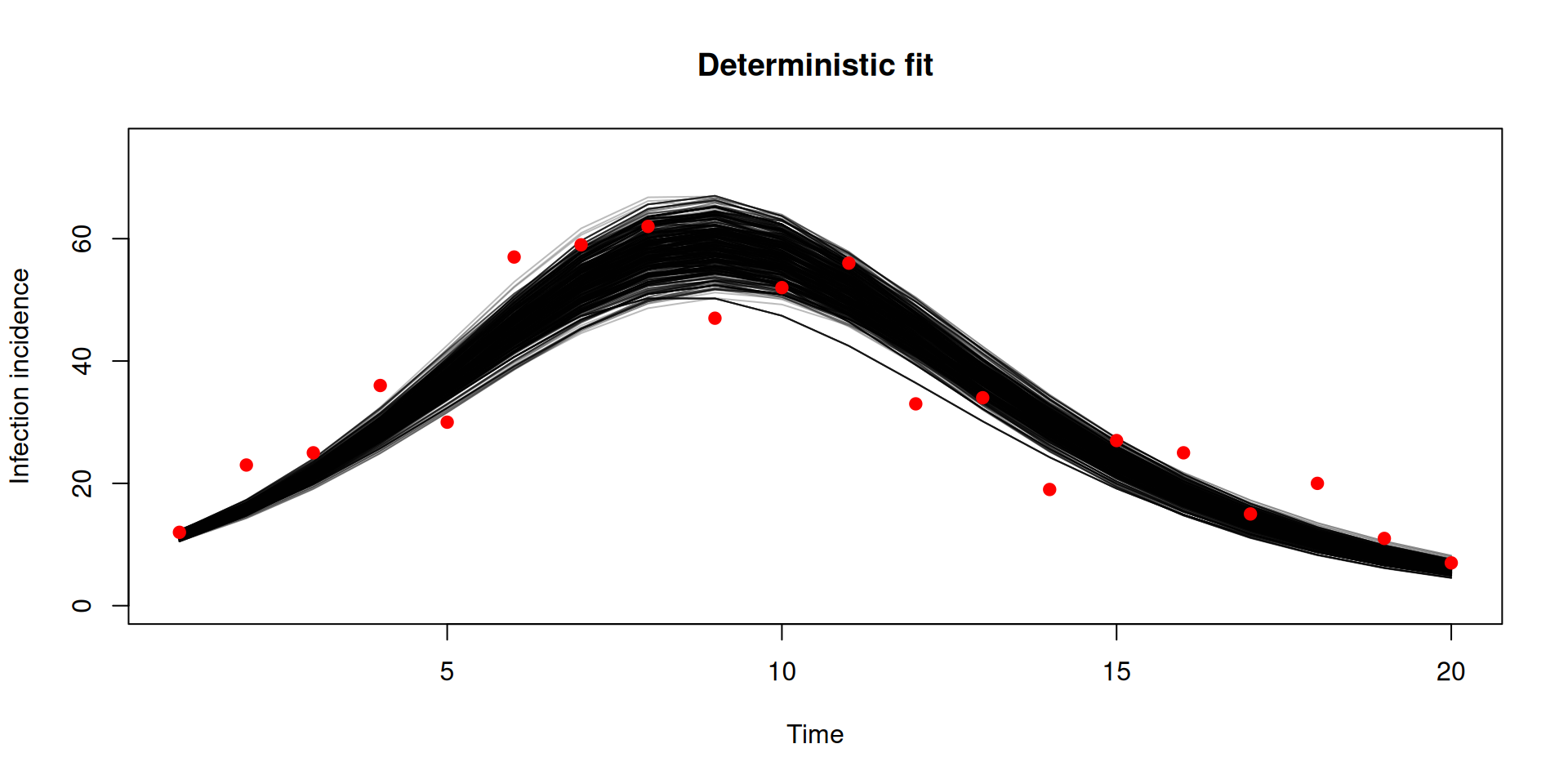
Stochastic v deterministic comparison
pars_stochastic <- array(samples$pars, c(2, 500))
pars_deterministic <- array(samples_det$pars, c(2, 500))
plot(pars_stochastic[1, ], pars_stochastic[2, ], ylab = "gamma", xlab = "beta",
pch = 19, col = "blue")
points(pars_deterministic[1, ], pars_deterministic[2, ], pch = 19, col = "red")
legend("bottomright", c("stochastic fit", "deterministic fit"), pch = c(19, 19),
col = c("blue", "red"))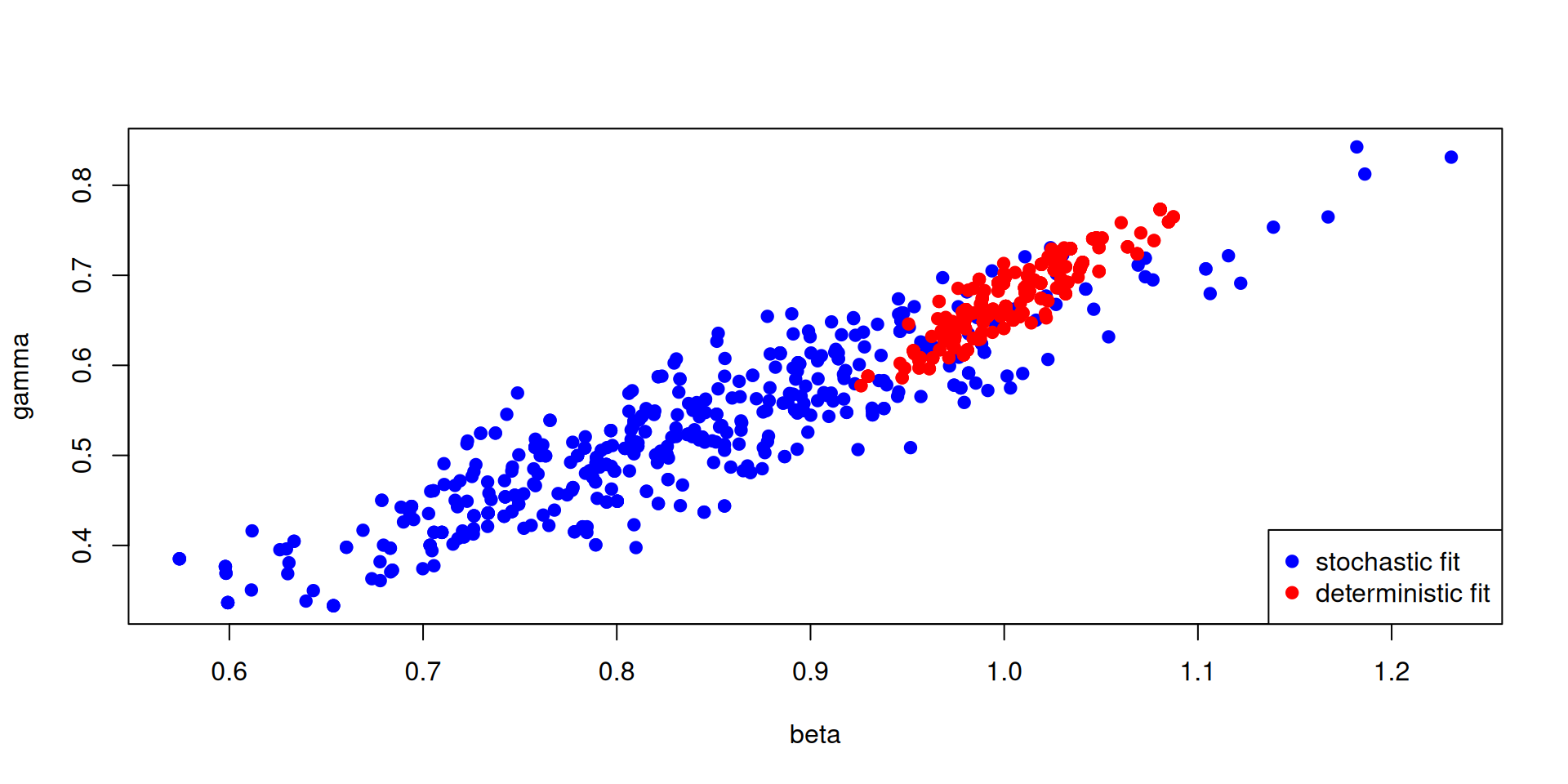
Fitting a time-varying parameter
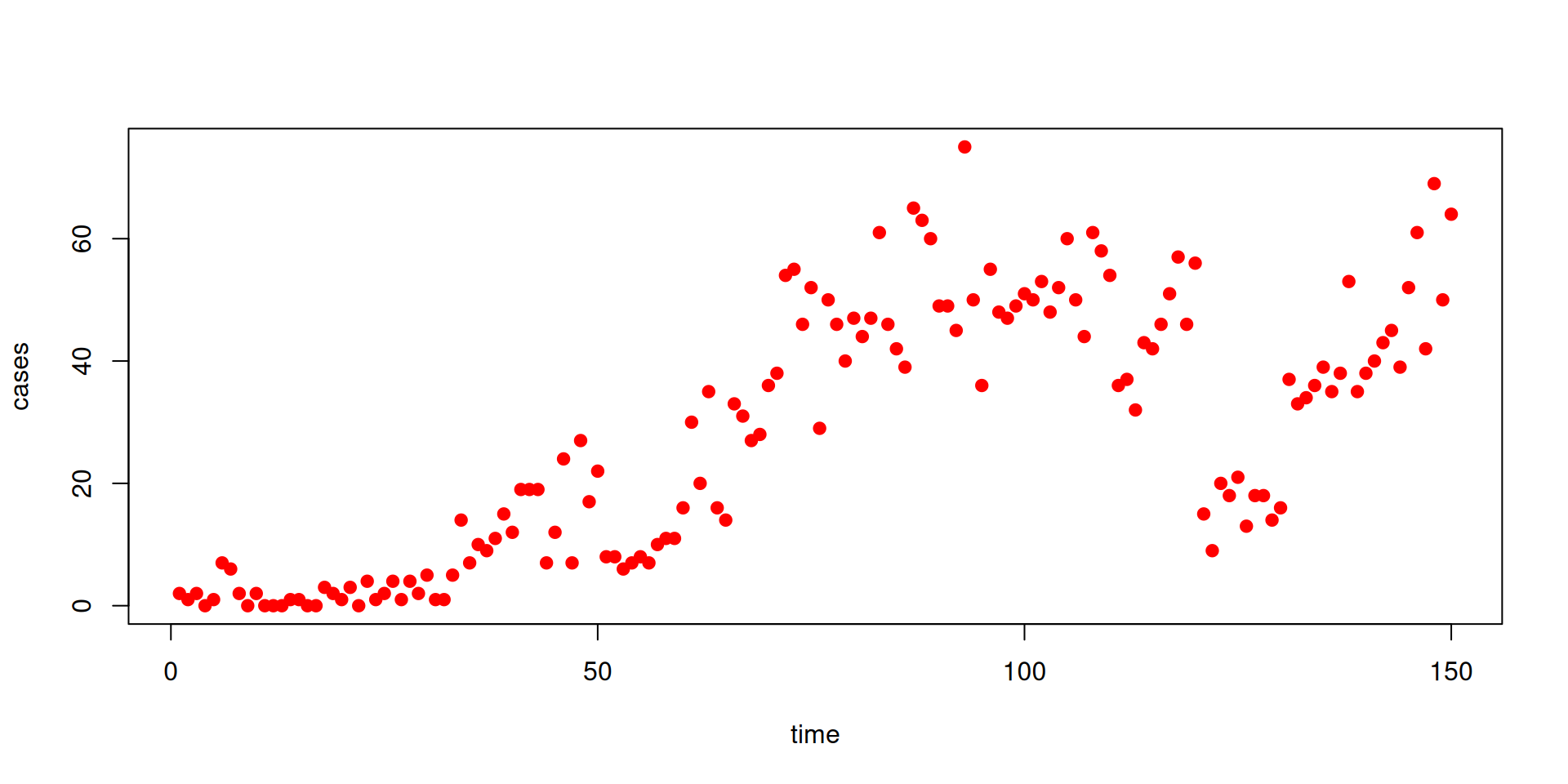
Let’s look at another dataset (the file can be downloaded here)
Fitting a time-varying parameter
Fitting a time-varying parameter
We’ll fit these data to a model with a piecewise constant beta using interpolate
sir <- odin({
update(S) <- S - n_SI
update(I) <- I + n_SI - n_IR
update(R) <- R + n_IR
update(incidence) <- incidence + n_SI
beta_t <- interpolate(beta_time, beta, "constant")
beta <- parameter()
beta_time <- parameter()
dim(beta, beta_time) <- parameter(rank = 1)
p_SI <- 1 - exp(-beta_t * I / N * dt)
p_IR <- 1 - exp(-gamma * dt)
n_SI <- Binomial(S, p_SI)
n_IR <- Binomial(I, p_IR)
initial(S) <- N - I0
initial(I) <- I0
initial(R) <- 0
initial(incidence, zero_every = 1) <- 0
N <- parameter(1000)
I0 <- parameter(10)
gamma <- parameter(0.1)
cases <- data()
cases ~ Poisson(incidence)
})Fitting a time-varying parameter
Let’s create our packer - we’ll fit gamma and 2 values for beta (with changepoints at time = 0 and time = 20)
Fitting a time-varying parameter
Let’s create our filter and likelihood model
and create a prior model. then combine to create our posterior model
Fitting a time-varying parameter
Combine to create our posterior model
posterior <- likelihood + prior
posterior
#>
#> ── <monty_model> ───────────────────────────────────────────────────────────────
#> ℹ Model has 3 parameters: 'gamma', 'beta[1]', and 'beta[2]'
#> ℹ This model:
#> • can be directly sampled from
#> • is stochastic
#> • has an observer
#> ℹ See `?monty_model()` for more informationCreate a sampler and then sample
Fitting a time-varying parameter
Fitting to vector/array data
Suppose we have some data on the daily incidence of cases by age group (the file can be downloaded here)
Let’s think of the cases data at each time point as a vector of length 2 (for the 2 age groups)
Fitting to vector/array data
We’ll fit these data to a model with age stratification
sir_age <- odin({
# Equations for transitions between compartments by age group
update(S[]) <- S[i] - n_SI[i]
update(I[]) <- I[i] + n_SI[i] - n_IR[i]
update(R[]) <- R[i] + n_IR[i]
update(incidence[]) <- incidence[i] + n_SI[i]
# Individual probabilities of transition:
p_SI[] <- 1 - exp(-lambda[i] * dt) # S to I
p_IR <- 1 - exp(-gamma * dt) # I to R
# Calculate force of infection
# age-structured contact matrix: m[i, j] is mean number
# of contacts an individual in group i has with an
# individual in group j per time unit
m <- parameter()
# here s_ij[i, j] gives the mean number of contacts an
# individual in group i will have with the currently
# infectious individuals of group j
s_ij[, ] <- m[i, j] * I[j]
# lambda[i] is the total force of infection on an
# individual in group i
lambda[] <- beta * sum(s_ij[i, ])
# Draws from binomial distributions for numbers
# changing between compartments:
n_SI[] <- Binomial(S[i], p_SI[i])
n_IR[] <- Binomial(I[i], p_IR)
initial(S[]) <- S0[i]
initial(I[]) <- I0[i]
initial(R[]) <- 0
initial(incidence[], zero_every = 1) <- 0
S0 <- parameter()
I0 <- parameter()
beta <- parameter(0.2)
gamma <- parameter(0.1)
n_age <- parameter()
dim(S, S0, n_SI, p_SI, incidence) <- n_age
dim(I, I0, n_IR) <- n_age
dim(R) <- n_age
dim(m, s_ij) <- c(n_age, n_age)
dim(lambda) <- n_age
## Likelihood
cases <- data()
dim(cases) <- n_age
cases[] ~ Poisson(incidence[i])
})Fitting to vector/array data
We are treating the data cases as a vector that relates to incidence from the model
- We can extend this to higher-dimensional arrays
- All datastreams that are arrays require
dim()equations
In the book: Data
Formatting array data in the data frame
Normally, you could setup a data.frame by writing:
Formatting array data in the data frame
Array data can be handled with a list-column.
For a list-column you pass in a list with the same number of elements as there are rows, with each list element being an array of the right size
Note we need to wrap with I()!
In the book: Data
Formatting array data in the data frame
Let’s convert our case columns to a single list-column (and then drop the separated case columns)
Fitting the age-stratified model
Let’s create our packer - we’ll fit beta and gamma and assume we know everything else
Fitting the age-stratified model
Let’s create our filter and likelihood model
and create a prior model, then combine to create our posterior model
Fitting the age-stratified model
Create a sampler and then sample
Fitting the age-stratified model
y <- dust_unpack_state(filter, samples$observations$trajectories)
incidence <- array(y$incidence, c(2, 100, 2000))
matplot(data_age$time, incidence[1, , ], type = "l", col = "#00000044", lty = 1,
xlab = "Time", ylab = "Incidence (children)")
cases_children <- vapply(data_age$cases, "[[", numeric(1), "cases_children")
points(data_age$time, cases_children, pch = 19, col = "red")
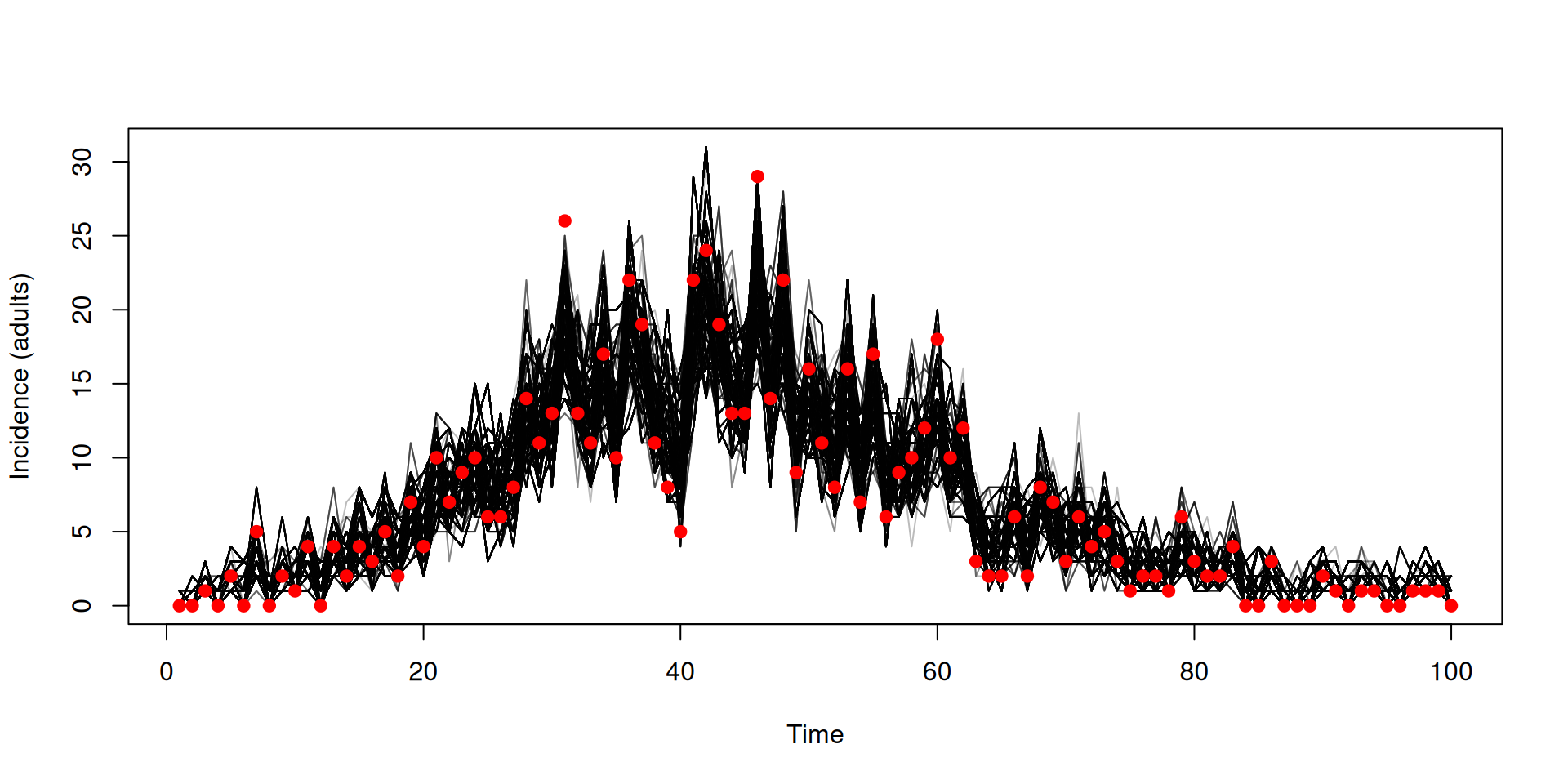
matplot(data_age$time, incidence[2, , ], type = "l", col = "#00000044", lty = 1,
xlab = "Time", ylab = "Incidence (adults)")
cases_adult <- vapply(data_age$cases, "[[", numeric(1), "cases_adult")
points(data_age$time, cases_adult, pch = 19, col = "red")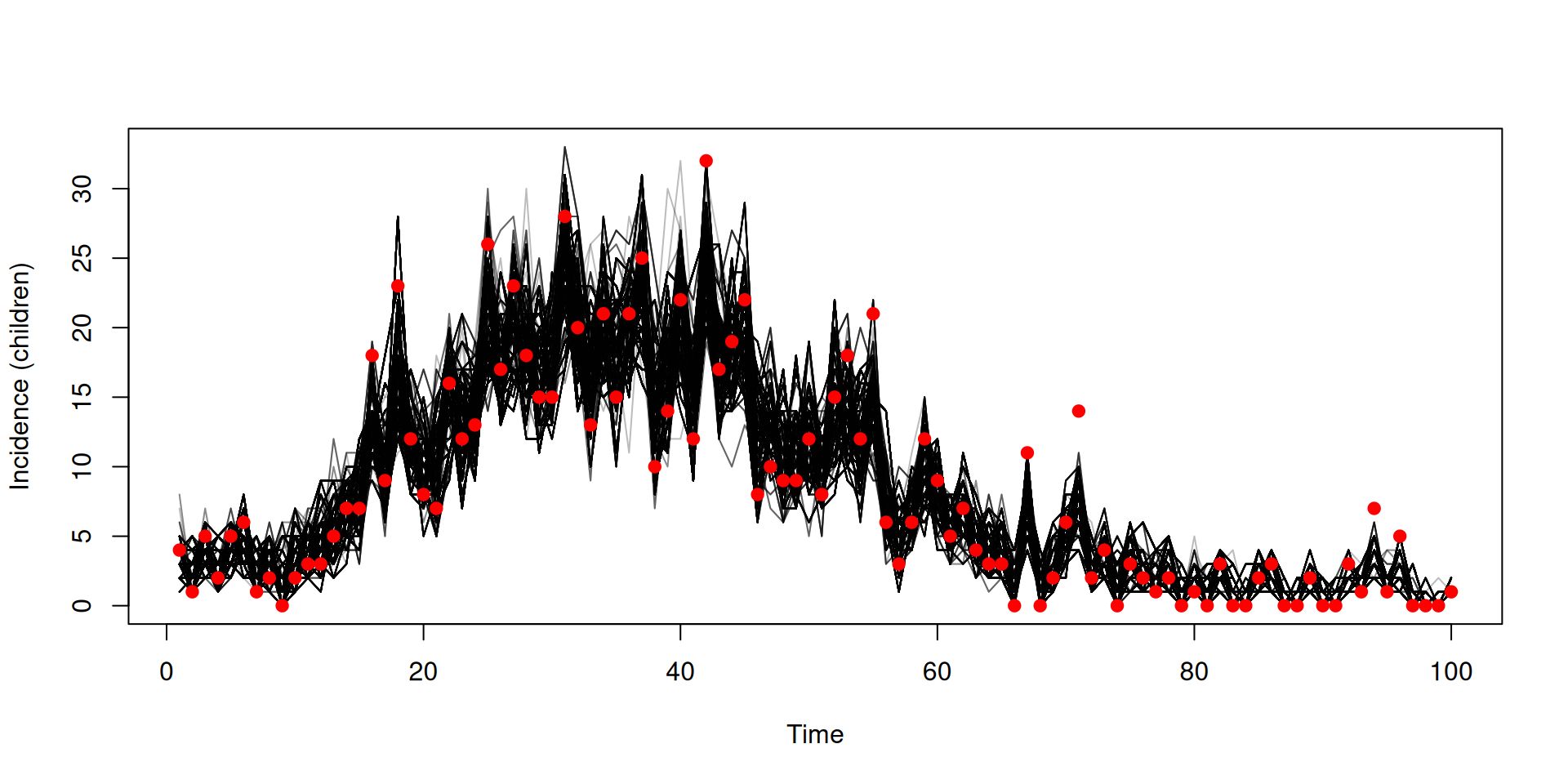
Projections and counterfactuals
Let’s use some new data (the file can be downloaded here)
In the book: Projections and counterfactuals
Projections and counterfactuals
We’ll fit the data to an SIS model incorporating schools opening/closing
sis <- odin({
update(S) <- S - n_SI + n_IS
update(I) <- I + n_SI - n_IS
update(incidence) <- incidence + n_SI
initial(S) <- N - I0
initial(I) <- I0
initial(incidence, zero_every = 1) <- 0
schools <- interpolate(schools_time, schools_open, "constant")
schools_time <- parameter()
schools_open <- parameter()
dim(schools_time, schools_open) <- parameter(rank = 1)
beta <- ((1 - schools) * (1 - schools_modifier) + schools) * beta0
p_SI <- 1 - exp(-beta * I / N * dt)
p_IS <- 1 - exp(-gamma * dt)
n_SI <- Binomial(S, p_SI)
n_IS <- Binomial(I, p_IS)
N <- parameter(1000)
I0 <- parameter(10)
beta0 <- parameter(0.2)
gamma <- parameter(0.1)
schools_modifier <- parameter(0.6)
cases <- data()
cases ~ Poisson(incidence)
})In the book: Projections and counterfactuals
Projections and counterfactuals
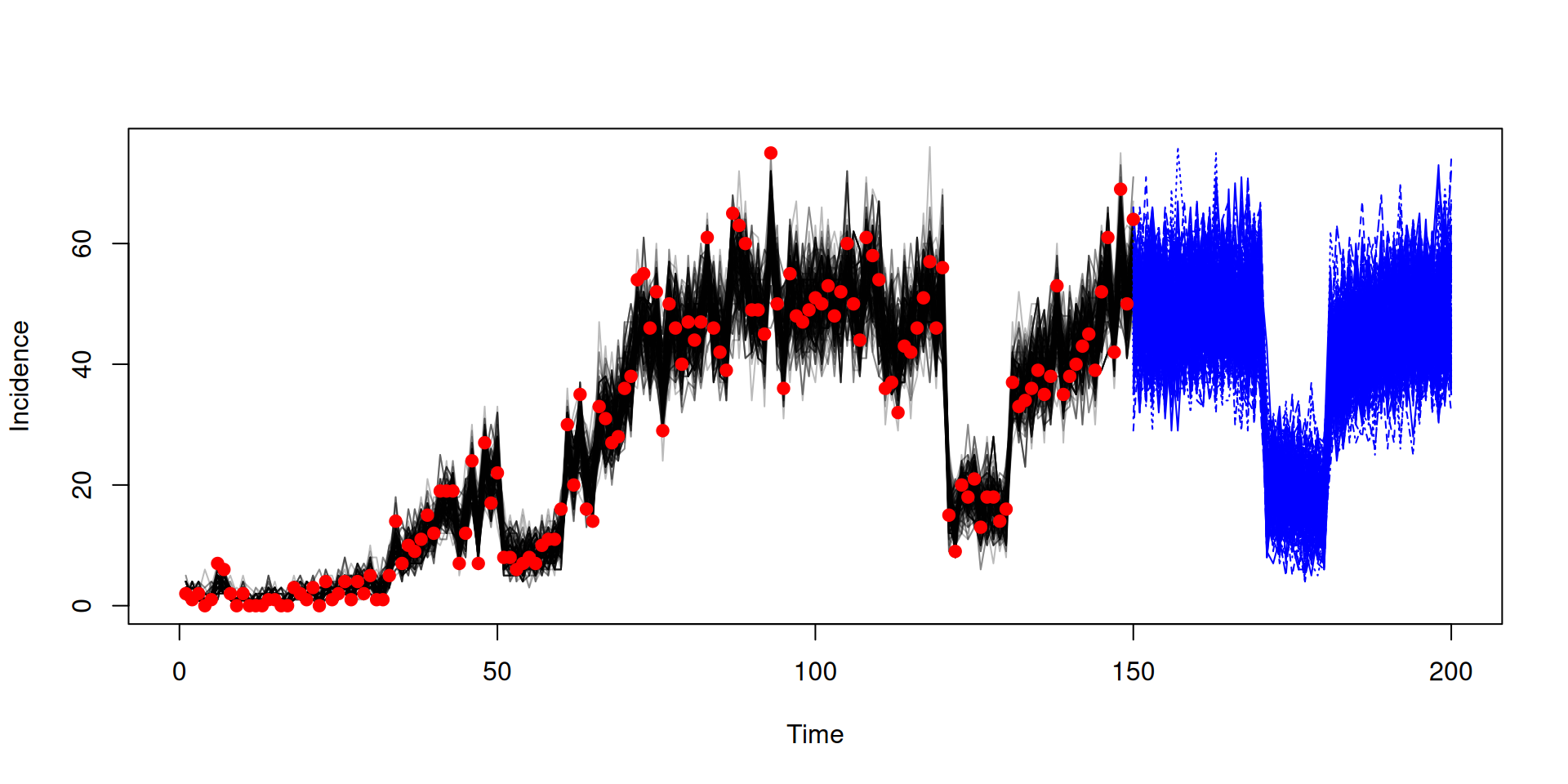
We will
- project forward from the end of the fits (day 150) to day 200
- run a counterfactual where the schools did not reopen on day 60, reopening on day 130
In the book: Projections and counterfactuals
Fitting to the SIS model
packer <- monty_packer(c("beta0", "gamma", "schools_modifier"),
fixed = list(schools_time = schools_time,
schools_open = schools_open))
data <- dust_filter_data(data, time = "time")
filter <- dust_filter_create(sis, time_start = 0, dt = 0.25,
data = data, n_particles = 200)
prior <- monty_dsl({
beta0 ~ Exponential(mean = 0.3)
gamma ~ Exponential(mean = 0.1)
schools_modifier ~ Uniform(0, 1)
})
vcv <- diag(c(2e-4, 2e-4, 4e-4))
sampler <- monty_sampler_random_walk(vcv)In the book: Projections and counterfactuals
Fitting to the SIS model
We want to save the end state, and a snapshot at day 60 (where the counterfactual will diverge)
likelihood <- dust_likelihood_monty(filter, packer, save_trajectories = TRUE,
save_state = TRUE, save_snapshots = 60)
posterior <- likelihood + prior
samples <- monty_sample(posterior, sampler, 500, initial = c(0.3, 0.1, 0.5),
n_chains = 4)
samples <- monty_samples_thin(samples, burnin = 100, thinning_factor = 8)In the book: Projections and counterfactuals
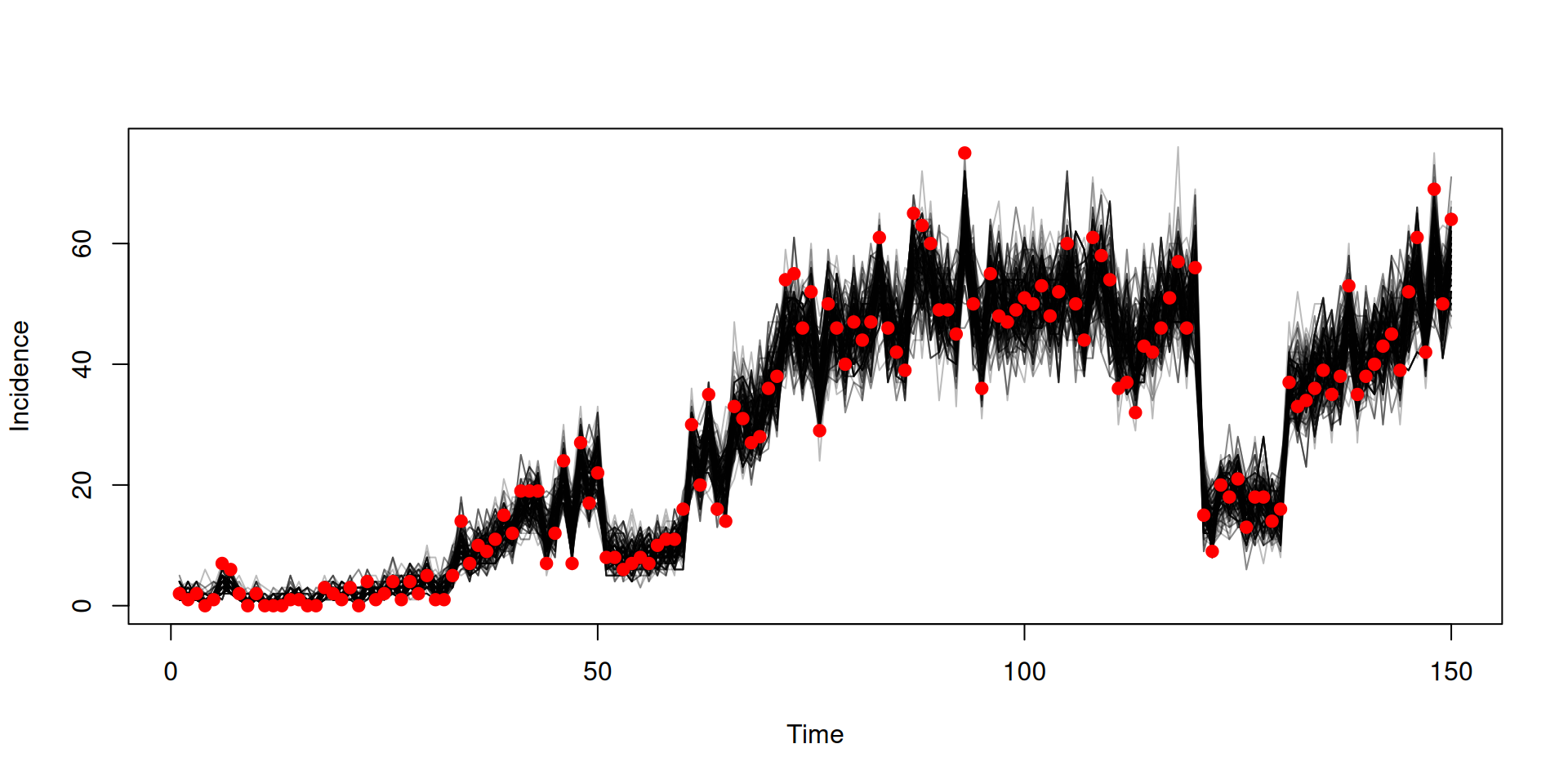
Fit to data
In the book: Projections and counterfactuals
Running projection using the end state
state <- array(samples$observations$state, c(3, 200))
pars <- array(samples$pars, c(3, 200))
pars <- lapply(seq_len(200), function(i) packer$unpack(pars[, i]))
sys <- dust_system_create(sis, pars, n_groups = length(pars), dt = 1)
dust_system_set_state(sys, state)
t <- seq(150, 200)
y <- dust_system_simulate(sys, t)
y <- dust_unpack_state(sys, y)In the book: Projections and counterfactuals
Running projection using the end state
In the book: Projections and counterfactuals
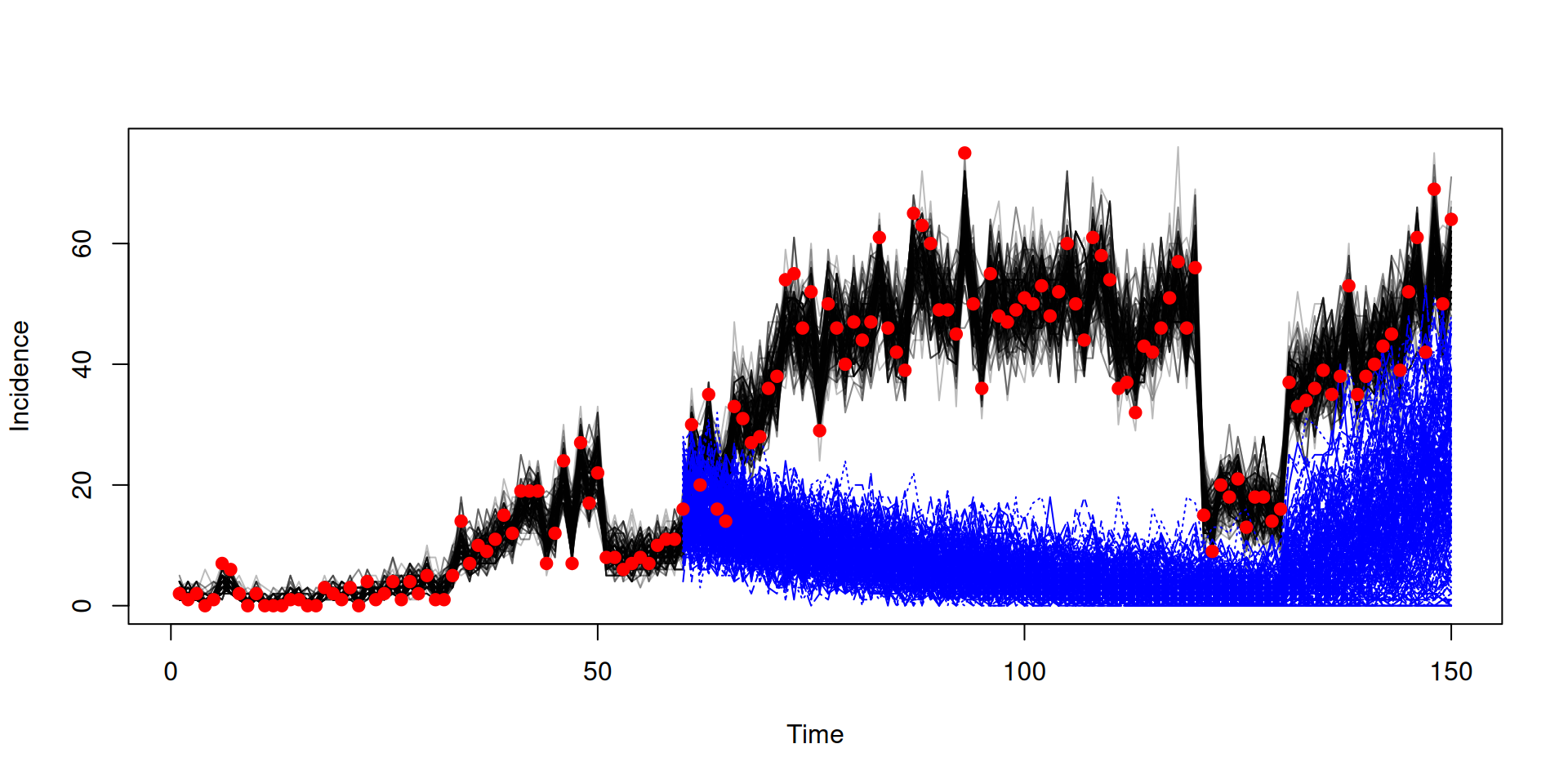
Running counterfactual using the snapshot
snapshot <- array(samples$observations$snapshots, c(3, 200))
pars <- array(samples$pars, c(3, 200))
f <- function(i) {
p <- packer$unpack(pars[, i])
p$schools_time <- c(0, 50, 130, 170, 180)
p$schools_open <- c(1, 0, 1, 0, 1)
p
}
pars <- lapply(seq_len(200), f)
sys <- dust_system_create(sis, pars, n_groups = length(pars), dt = 1)
dust_system_set_state(sys, snapshot)
t <- seq(60, 150)
y <- dust_system_simulate(sys, t)
y <- dust_unpack_state(sys, y)In the book: Projections and counterfactuals
Running counterfactual using the snapshot
In the book: Projections and counterfactuals
Next steps
Try different samplers
- adaptive sampler (for deterministic models)
Some under development to support odin models
- Hamiltonian Monte Carlo (HMC)
- parallel tempering
- nested sampler for grouped models